Host cell poly(ADP-ribose) glycohydrolase is crucial for Trypanosoma cruzi infection cycle
- PMID: 23776710
- PMCID: PMC3680488
- DOI: 10.1371/journal.pone.0067356
Host cell poly(ADP-ribose) glycohydrolase is crucial for Trypanosoma cruzi infection cycle
Abstract
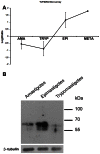
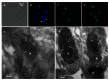
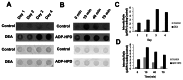
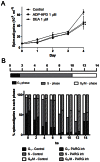
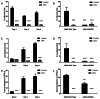
Trypanosoma cruzi, etiological agent of Chagas' disease, has a complex life cycle which involves the invasion of mammalian host cells, differentiation and intracellular replication. Here we report the first insights into the biological role of a poly(ADP-ribose) glycohydrolase in a trypanosomatid (TcPARG). In silico analysis of the TcPARG gene pointed out the conservation of key residues involved in the catalytic process and, by Western blot, we demonstrated that it is expressed in a life stage-dependant manner. Indirect immunofluorescence assays and electron microscopy using an anti-TcPARG antibody showed that this enzyme is localized in the nucleus independently of the presence of DNA damage or cell cycle stage. The addition of poly(ADP-ribose) glycohydrolase inhibitors ADP-HPD (adenosine diphosphate (hydroxymethyl) pyrrolidinediol) or DEA (6,9-diamino-2-ethoxyacridine lactate monohydrate) to the culture media, both at a 1 µM concentration, reduced in vitro epimastigote growth by 35% and 37% respectively, when compared to control cultures. We also showed that ADP-HPD 1 µM can lead to an alteration in the progression of the cell cycle in hydroxyurea synchronized cultures of T. cruzi epimastigotes. Outstandingly, here we demonstrate that the lack of poly(ADP-ribose) glycohydrolase activity in Vero and A549 host cells, achieved by chemical inhibition or iRNA, produces the reduction of the percentage of infected cells as well as the number of amastigotes per cell and trypomastigotes released, leading to a nearly complete abrogation of the infection process. We conclude that both, T. cruzi and the host, poly(ADP-ribose) glycohydrolase activities are important players in the life cycle of Trypanosoma cruzi, emerging as a promising therapeutic target for the treatment of Chagas' disease.
Conflict of interest statement
Figures

Similar articles
-
Syntheses of photoactive analogues of adenosine diphosphate (hydroxymethyl)pyrrolidinediol and photoaffinity labeling of poly(ADP-ribose) glycohydrolase.Biochemistry. 1998 May 26;37(21):7801-12. doi: 10.1021/bi9730386. Biochemistry. 1998. PMID: 9601041
-
Mechanism of inhibition of poly(ADP-ribose) glycohydrolase by adenosine diphosphate (hydroxymethyl)pyrrolidinediol.J Med Chem. 1995 Oct 13;38(21):4332-6. doi: 10.1021/jm00021a023. J Med Chem. 1995. PMID: 7473561
-
Specific inhibition of poly(ADP-ribose) glycohydrolase by adenosine diphosphate (hydroxymethyl)pyrrolidinediol.J Med Chem. 1995 Jan 20;38(2):389-93. doi: 10.1021/jm00002a021. J Med Chem. 1995. PMID: 7830282
-
Host cell actin remodeling in response to Trypanosoma cruzi: trypomastigote versus amastigote entry.Subcell Biochem. 2008;47:101-9. doi: 10.1007/978-0-387-78267-6_8. Subcell Biochem. 2008. PMID: 18512345 Review.
-
Importance of poly(ADP-ribose) glycohydrolase in the control of poly(ADP-ribose) metabolism.Exp Cell Res. 2001 Aug 1;268(1):7-13. doi: 10.1006/excr.2001.5263. Exp Cell Res. 2001. PMID: 11461113 Review.
Cited by
-
Proximal ADP-ribose Hydrolysis in Trypanosomatids is Catalyzed by a Macrodomain.Sci Rep. 2016 Apr 11;6:24213. doi: 10.1038/srep24213. Sci Rep. 2016. PMID: 27064071 Free PMC article.
-
Disrupted ADP-ribose metabolism with nuclear Poly (ADP-ribose) accumulation leads to different cell death pathways in presence of hydrogen peroxide in procyclic Trypanosoma brucei.Parasit Vectors. 2016 Mar 23;9:173. doi: 10.1186/s13071-016-1461-1. Parasit Vectors. 2016. PMID: 27007296 Free PMC article.
-
Roles of ADP-Ribosylation during Infection Establishment by Trypanosomatidae Parasites.Pathogens. 2023 May 12;12(5):708. doi: 10.3390/pathogens12050708. Pathogens. 2023. PMID: 37242378 Free PMC article. Review.
-
Tankyrase inhibitors hinder Trypanosoma cruzi infection by altering host-cell signalling pathways.Parasitology. 2021 Nov;148(13):1680-1690. doi: 10.1017/S0031182021001402. Epub 2021 Aug 12. Parasitology. 2021. PMID: 35060470 Free PMC article.
-
Poly(ADP-Ribose) Polymerases in Host-Pathogen Interactions, Inflammation, and Immunity.Microbiol Mol Biol Rev. 2018 Dec 19;83(1):e00038-18. doi: 10.1128/MMBR.00038-18. Print 2019 Mar. Microbiol Mol Biol Rev. 2018. PMID: 30567936 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

