Pbx and Prdm1a transcription factors differentially regulate subsets of the fast skeletal muscle program in zebrafish
- PMID: 23789105
- PMCID: PMC3683157
- DOI: 10.1242/bio.20133921
Pbx and Prdm1a transcription factors differentially regulate subsets of the fast skeletal muscle program in zebrafish
Abstract
The basic helix-loop-helix factor Myod initiates skeletal muscle differentiation by directly and sequentially activating sets of muscle differentiation genes, including those encoding muscle contractile proteins. We hypothesize that Pbx homeodomain proteins direct Myod to a subset of its transcriptional targets, in particular fast-twitch muscle differentiation genes, thereby regulating the competence of muscle precursor cells to differentiate. We have previously shown that Pbx proteins bind with Myod on the promoter of the zebrafish fast muscle gene mylpfa and that Pbx proteins are required for Myod to activate mylpfa expression and the fast-twitch muscle-specific differentiation program in zebrafish embryos. Here we have investigated the interactions of Pbx with another muscle fiber-type regulator, Prdm1a, a SET-domain DNA-binding factor that directly represses mylpfa expression and fast muscle differentiation. The prdm1a mutant phenotype, early and increased fast muscle differentiation, is the opposite of the Pbx-null phenotype, delayed and reduced fast muscle differentiation. To determine whether Pbx and Prdm1a have opposing activities on a common set of genes, we used RNA-seq analysis to globally assess gene expression in zebrafish embryos with single- and double-losses-of-function for Pbx and Prdm1a. We find that the levels of expression of certain fast muscle genes are increased or approximately wild type in pbx2/4-MO;prdm1a-/- embryos, suggesting that Pbx activity normally counters the repressive action of Prdm1a for a subset of the fast muscle program. However, other fast muscle genes require Pbx but are not regulated by Prdm1a. Thus, our findings reveal that subsets of the fast muscle program are differentially regulated by Pbx and Prdm1a. Our findings provide an example of how Pbx homeodomain proteins act in a balance with other transcription factors to regulate subsets of a cellular differentiation program.
Keywords: Fiber-type differentiation; Pbx; Prdm1; Skeletal muscle; Zebrafish.
Conflict of interest statement
Figures


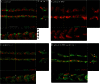
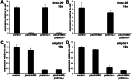

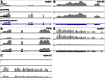
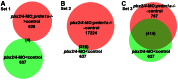


Similar articles
-
Pbx homeodomain proteins direct Myod activity to promote fast-muscle differentiation.Development. 2007 Sep;134(18):3371-82. doi: 10.1242/dev.003905. Epub 2007 Aug 15. Development. 2007. PMID: 17699609
-
Pbx acts with Hand2 in early myocardial differentiation.Dev Biol. 2009 Sep 15;333(2):409-18. doi: 10.1016/j.ydbio.2009.07.004. Epub 2009 Jul 14. Dev Biol. 2009. PMID: 19607825 Free PMC article.
-
Pbx homeodomain proteins pattern both the zebrafish retina and tectum.BMC Dev Biol. 2007 Jul 16;7:85. doi: 10.1186/1471-213X-7-85. BMC Dev Biol. 2007. PMID: 17634100 Free PMC article.
-
Control of muscle fibre-type diversity during embryonic development: the zebrafish paradigm.Mech Dev. 2013 Sep-Oct;130(9-10):447-57. doi: 10.1016/j.mod.2013.06.001. Epub 2013 Jun 28. Mech Dev. 2013. PMID: 23811405 Review.
-
Regulating a master regulator: establishing tissue-specific gene expression in skeletal muscle.Epigenetics. 2010 Nov-Dec;5(8):691-5. doi: 10.4161/epi.5.8.13045. Epub 2010 Nov 1. Epigenetics. 2010. PMID: 20716948 Free PMC article. Review.
Cited by
-
'Building a perfect body': control of vertebrate organogenesis by PBX-dependent regulatory networks.Genes Dev. 2019 Mar 1;33(5-6):258-275. doi: 10.1101/gad.318774.118. Genes Dev. 2019. PMID: 30824532 Free PMC article. Review.
-
Pbx4 is Required for the Temporal Onset of Zebrafish Myocardial Differentiation.J Dev Biol. 2015;3(4):93-111. doi: 10.3390/jdb3040093. J Dev Biol. 2015. PMID: 26770887 Free PMC article.
-
Contrasting roles for MyoD in organizing myogenic promoter structures during embryonic skeletal muscle development.Dev Dyn. 2015 Jan;244(1):43-55. doi: 10.1002/dvdy.24217. Epub 2014 Nov 3. Dev Dyn. 2015. PMID: 25329411 Free PMC article.
-
Cell fusion is differentially regulated in zebrafish post-embryonic slow and fast muscle.Dev Biol. 2020 Jun 1;462(1):85-100. doi: 10.1016/j.ydbio.2020.03.005. Epub 2020 Mar 10. Dev Biol. 2020. PMID: 32165147 Free PMC article.
-
Myogenin promotes myocyte fusion to balance fibre number and size.Nat Commun. 2018 Oct 12;9(1):4232. doi: 10.1038/s41467-018-06583-6. Nat Commun. 2018. PMID: 30315160 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

