RNA-seq in the tetraploid Xenopus laevis enables genome-wide insight in a classic developmental biology model organism
- PMID: 23792920
- PMCID: PMC3884041
- DOI: 10.1016/j.ymeth.2013.06.009
RNA-seq in the tetraploid Xenopus laevis enables genome-wide insight in a classic developmental biology model organism
Abstract
Advances in sequencing technology have significantly advanced the landscape of developmental biology research. The dissection of genetic networks in model and non-model organisms has been greatly enhanced with high-throughput sequencing technologies. RNA-seq has revolutionized the ability to perform developmental biology research in organisms without a published genome sequence. Here, we describe a protocol for developmental biologists to perform RNA-seq on dissected tissue or whole embryos. We start with the isolation of RNA and generation of sequencing libraries. We further show how to interpret and analyze the large amount of sequencing data that is generated in RNA-seq. We explore the abilities to examine differential expression, gene duplication, transcript assembly, alternative splicing and SNP discovery. For the purposes of this article, we use Xenopus laevis as the model organism to discuss uses of RNA-seq in an organism without a fully annotated genome sequence.
Keywords: Differential expression; RNA-seq; Xenopus.
Copyright © 2013. Published by Elsevier Inc.
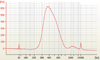
Figures



References
-
- Nusslein-Volhard C, Wieschaus E. Nature. 1980;287:795–801. - PubMed
-
- Haffter P, Granato M, Brand M, Mullins MC, Hammerschmidt M, Kane DA, Odenthal J, van Eeden FJ, Jiang YJ, Heisenberg CP, Kelsh RN, Furutani-Seiki M, Vogelsang E, Beuchle D, Schach U, Fabian C, Nusslein-Volhard C. Development. 1996;123:1–36. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

