Mutational analysis of Sse1 (Hsp110) suggests an integral role for this chaperone in yeast prion propagation in vivo
- PMID: 23797105
- PMCID: PMC3737180
- DOI: 10.1534/g3.113.007112
Mutational analysis of Sse1 (Hsp110) suggests an integral role for this chaperone in yeast prion propagation in vivo
Abstract
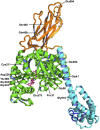
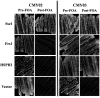
The yeast Hsp110 chaperone Sse1 is a conserved protein that is a noncanonical member of the Hsp70 protein superfamily. Sse1 influences the cellular response to heat stress and has also been implicated in playing a role in the propagation of prions in yeast. Sse1 can seemingly exert its effects in vivo through direct or indirect actions by influencing the nucleotide exchange activity of canonical cytosolic Hsp70s. Using a genetic screen based on the inability to propagate the yeast [PSI(+)] prion, we have identified 13 new Sse1 mutants that are predicted to alter chaperone function through a variety of different mechanisms. Not only are these new Sse1 mutants altered in the ability to propagate and cure yeast prions but also to varying degrees in the ability to grow at elevated temperatures. The expression levels of chaperone proteins known to influence yeast prion propagation are unaltered in the Sse1 mutants, suggesting that the observed phenotypic effects are caused by direct functional alterations in these mutants. Mapping the location of the mutants onto the Sse1 crystal structure suggests that more than one functional alteration in Sse1 may result in changes in prion propagation and ability to function at elevated temperatures. All Sse1 mutants isolated provide essential functions in the cell under normal growth conditions, further demonstrating that essential chaperone functions in vivo can to some degree at least be detached from those related to propagation of prions. Our results suggest that Sse1 can influence prion propagation through a variety of different mechanisms.
Keywords: Hsp110; Hsp70; Saccharomyces cerevisiae; Sse1; chaperone; nucleotide exchange factor; prion.
Figures




Similar articles
-
Nucleotide exchange factors for Hsp70s are required for [URE3] prion propagation in Saccharomyces cerevisiae.Mol Biol Cell. 2007 Jun;18(6):2149-54. doi: 10.1091/mbc.e07-02-0128. Epub 2007 Mar 28. Mol Biol Cell. 2007. PMID: 17392510 Free PMC article.
-
The role of Sse1 in the de novo formation and variant determination of the [PSI+] prion.Genetics. 2007 Nov;177(3):1583-93. doi: 10.1534/genetics.107.077982. Genetics. 2007. PMID: 18039878 Free PMC article.
-
Characterization of Hsp70 binding and nucleotide exchange by the yeast Hsp110 chaperone Sse1.Biochemistry. 2006 Dec 19;45(50):15075-84. doi: 10.1021/bi061279k. Biochemistry. 2006. PMID: 17154545 Free PMC article.
-
Impact of Amyloid Polymorphism on Prion-Chaperone Interactions in Yeast.Viruses. 2019 Apr 16;11(4):349. doi: 10.3390/v11040349. Viruses. 2019. PMID: 30995727 Free PMC article. Review.
-
Influence of Hsp70s and their regulators on yeast prion propagation.Prion. 2009 Apr-Jun;3(2):65-73. doi: 10.4161/pri.3.2.9134. Epub 2009 Apr 29. Prion. 2009. PMID: 19556854 Free PMC article. Review.
Cited by
-
Protein rescue from aggregates by powerful molecular chaperone machines.Nat Rev Mol Cell Biol. 2013 Oct;14(10):617-29. doi: 10.1038/nrm3660. Nat Rev Mol Cell Biol. 2013. PMID: 24061228 Review.
-
Hsp104 disaggregase at normal levels cures many [PSI+] prion variants in a process promoted by Sti1p, Hsp90, and Sis1p.Proc Natl Acad Sci U S A. 2017 May 23;114(21):E4193-E4202. doi: 10.1073/pnas.1704016114. Epub 2017 May 8. Proc Natl Acad Sci U S A. 2017. PMID: 28484020 Free PMC article.
-
Purification and biochemical characterization of Msi3, an essential Hsp110 molecular chaperone in Candida albicans.Cell Stress Chaperones. 2021 Jul;26(4):695-704. doi: 10.1007/s12192-021-01213-5. Epub 2021 May 28. Cell Stress Chaperones. 2021. PMID: 34047887 Free PMC article.
-
Dancing through Life: Molecular Dynamics Simulations and Network-Centric Modeling of Allosteric Mechanisms in Hsp70 and Hsp110 Chaperone Proteins.PLoS One. 2015 Nov 30;10(11):e0143752. doi: 10.1371/journal.pone.0143752. eCollection 2015. PLoS One. 2015. PMID: 26619280 Free PMC article.
-
A first-in-class inhibitor of Hsp110 molecular chaperones of pathogenic fungi.Nat Commun. 2023 May 12;14(1):2745. doi: 10.1038/s41467-023-38220-2. Nat Commun. 2023. PMID: 37173314 Free PMC article.
References
-
- Bach S., Talarek N., Andrieu T., Vierfond J. M., Mettey Y., et al. , 2003. Isolation of drugs active against mammalian prions using a yeast-based screening assay. Nat. Biotechnol. 21: 1075–1081 - PubMed
-
- Chemical Computing Group Inc., 2009. Molecular Operating Environment, Version 2009.10. Chemical Computing Group, Inc., Montreal, QC, Canada
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
