Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force
- PMID: 23798731
- PMCID: PMC3691458
- DOI: 10.1083/jcb.201210152
Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force
Abstract
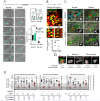
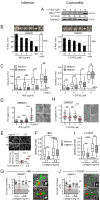
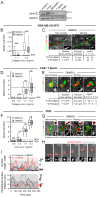
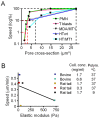
Cell migration through 3D tissue depends on a physicochemical balance between cell deformability and physical tissue constraints. Migration rates are further governed by the capacity to degrade ECM by proteolytic enzymes, particularly matrix metalloproteinases (MMPs), and integrin- and actomyosin-mediated mechanocoupling. Yet, how these parameters cooperate when space is confined remains unclear. Using MMP-degradable collagen lattices or nondegradable substrates of varying porosity, we quantitatively identify the limits of cell migration by physical arrest. MMP-independent migration declined as linear function of pore size and with deformation of the nucleus, with arrest reached at 10% of the nuclear cross section (tumor cells, 7 µm²; T cells, 4 µm²; neutrophils, 2 µm²). Residual migration under space restriction strongly depended upon MMP-dependent ECM cleavage by enlarging matrix pore diameters, and integrin- and actomyosin-dependent force generation, which jointly propelled the nucleus. The limits of interstitial cell migration thus depend upon scaffold porosity and deformation of the nucleus, with pericellular collagenolysis and mechanocoupling as modulators.
Figures




Comment in
-
Cell migration: Feeling the squeeze.Nat Rev Mol Cell Biol. 2013 Aug;14(8):462. doi: 10.1038/nrm3627. Epub 2013 Jul 10. Nat Rev Mol Cell Biol. 2013. PMID: 23839577 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

