Mitochondrial network morphology: building an integrative, geometrical view
- PMID: 23800141
- PMCID: PMC3691739
- DOI: 10.1186/1741-7007-11-71
Mitochondrial network morphology: building an integrative, geometrical view
Abstract
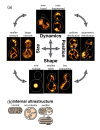
The morphology of mitochondrial networks is complex and highly varied, yet vital to cell function. The first step toward an integrative understanding of how mitochondrial morphology is generated and regulated is to define the interdependent geometrical features and their dynamics that together generate the morphology of a mitochondrial network within a cell. Distinct aspects of the size, shape, position, and dynamics of mitochondrial networks are described and examples of how these features depend on one another discussed.
Figures


Comment in
-
AI spots cell structures that humans can't.Nature. 2021 Apr;592(7852):154-155. doi: 10.1038/d41586-021-00812-7. Nature. 2021. PMID: 33782599 No abstract available.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

