Modeling regulatory cascades using Artificial Neural Networks: the case of transcriptional regulatory networks shaped during the yeast stress response
- PMID: 23802010
- PMCID: PMC3687159
- DOI: 10.3389/fgene.2013.00110
Modeling regulatory cascades using Artificial Neural Networks: the case of transcriptional regulatory networks shaped during the yeast stress response
Abstract
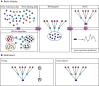
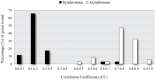
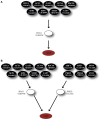
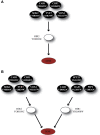
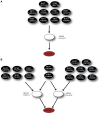
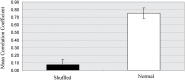
Over the last decade, numerous computational methods have been developed in order to infer and model biological networks. Transcriptional networks in particular have attracted significant attention due to their critical role in cell survival. The majority of network inference methods use genome-wide experimental data to search for modules of genes with coherent expression profiles and common regulators, often ignoring the multi-layer structure of transcriptional cascades. Modeling methodologies on the other hand assume a given network structure and vary significantly in their algorithmic approach, ranging from over-simplified representations (e.g., Boolean networks) to detailed -but computationally expensive-network simulations (e.g., with differential equations). In this work we use Artificial Neural Networks (ANNs) to model transcriptional regulatory cascades that emerge during the stress response in Saccharomyces cerevisiae and extend in three layers. We confine the structure of the ANNs to match the structure of the biological networks as determined by gene expression, DNA-protein interaction and experimental evidence provided in publicly available databases. Trained ANNs are able to predict the expression profile of 11 target genes across multiple experimental conditions with a correlation coefficient >0.7. When time-dependent interactions between upstream transcription factors (TFs) and their indirect targets are also included in the ANNs, accurate predictions are achieved for 30/34 target genes. Moreover, heterodimer formation is taken into account. We show that ANNs can be used to (1) accurately predict the expression of downstream genes in a 3-layer transcriptional cascade based on the expression of their indirect regulators and (2) infer the condition- and time-dependent activity of various TFs as well as during heterodimer formation. We show that a three-layer regulatory cascade whose structure is determined by co-expressed gene modules and their regulators can successfully be modeled using ANNs with a similar configuration.
Keywords: Artificial Neural Networks; asynchronous regulation; heterodimers; three layers regulatory cascades; transcriptional regulatory networks; yeast stress response.
Figures





Similar articles
-
Network motif-based identification of transcription factor-target gene relationships by integrating multi-source biological data.BMC Bioinformatics. 2008 Apr 21;9:203. doi: 10.1186/1471-2105-9-203. BMC Bioinformatics. 2008. PMID: 18426580 Free PMC article.
-
MICRAT: a novel algorithm for inferring gene regulatory networks using time series gene expression data.BMC Syst Biol. 2018 Dec 14;12(Suppl 7):115. doi: 10.1186/s12918-018-0635-1. BMC Syst Biol. 2018. PMID: 30547796 Free PMC article.
-
Improved recovery of cell-cycle gene expression in Saccharomyces cerevisiae from regulatory interactions in multiple omics data.BMC Genomics. 2020 Feb 13;21(1):159. doi: 10.1186/s12864-020-6554-8. BMC Genomics. 2020. PMID: 32054475 Free PMC article.
-
Mapping yeast transcriptional networks.Genetics. 2013 Sep;195(1):9-36. doi: 10.1534/genetics.113.153262. Genetics. 2013. PMID: 24018767 Free PMC article. Review.
-
Gene Regulatory Network Inference: Connecting Plant Biology and Mathematical Modeling.Front Genet. 2020 May 25;11:457. doi: 10.3389/fgene.2020.00457. eCollection 2020. Front Genet. 2020. PMID: 32547596 Free PMC article. Review.
Cited by
-
Stress tolerance enhancement via SPT15 base editing in Saccharomyces cerevisiae.Biotechnol Biofuels. 2021 Jul 6;14(1):155. doi: 10.1186/s13068-021-02005-w. Biotechnol Biofuels. 2021. PMID: 34229745 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

