The tRNA recognition mechanism of the minimalist SPOUT methyltransferase, TrmL
- PMID: 23804755
- PMCID: PMC3763551
- DOI: 10.1093/nar/gkt568
The tRNA recognition mechanism of the minimalist SPOUT methyltransferase, TrmL
Abstract
Unlike other transfer RNAs (tRNA)-modifying enzymes from the SPOUT methyltransferase superfamily, the tRNA (Um34/Cm34) methyltransferase TrmL lacks the usual extension domain for tRNA binding and consists only of a SPOUT domain. Both the catalytic and tRNA recognition mechanisms of this enzyme remain elusive. By using tRNAs purified from an Escherichia coli strain with the TrmL gene deleted, we found that TrmL can independently catalyze the methyl transfer from S-adenosyl-L-methionine to and isoacceptors without the involvement of other tRNA-binding proteins. We have solved the crystal structures of TrmL in apo form and in complex with S-adenosyl-homocysteine and identified the cofactor binding site and a possible active site. Methyltransferase activity and tRNA-binding affinity of TrmL mutants were measured to identify residues important for tRNA binding of TrmL. Our results suggest that TrmL functions as a homodimer by using the conserved C-terminal half of the SPOUT domain for catalysis, whereas residues from the less-conserved N-terminal half of the other subunit participate in tRNA recognition.
Figures

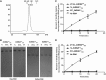
 s and
s and  s, respectively.
s, respectively.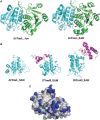
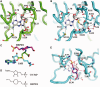


 analyzed by the gel mobility shift assay. (B) The methyltransferase activities of the various EcTrmL mutants.
analyzed by the gel mobility shift assay. (B) The methyltransferase activities of the various EcTrmL mutants.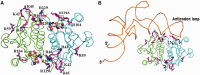
 is shown in brown.
is shown in brown.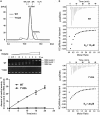
 was analyzed by the gel mobility shift assay. (C) The methyltransferase activity of Y142A. (D) The binding affinity for SAH by ITC.
was analyzed by the gel mobility shift assay. (C) The methyltransferase activity of Y142A. (D) The binding affinity for SAH by ITC.Similar articles
-
Identification of determinants for tRNA substrate recognition by Escherichia coli C/U34 2'-O-methyltransferase.RNA Biol. 2015;12(8):900-11. doi: 10.1080/15476286.2015.1050576. RNA Biol. 2015. PMID: 26106808 Free PMC article.
-
Structural and biochemical insights into the 2'-O-methylation of pyrimidines 34 in tRNA.FEBS J. 2017 Jul;284(14):2251-2263. doi: 10.1111/febs.14120. Epub 2017 Jun 18. FEBS J. 2017. PMID: 28544464
-
tRNA recognition by a bacterial tRNA Xm32 modification enzyme from the SPOUT methyltransferase superfamily.Nucleic Acids Res. 2015 Sep 3;43(15):7489-503. doi: 10.1093/nar/gkv745. Epub 2015 Jul 21. Nucleic Acids Res. 2015. PMID: 26202969 Free PMC article.
-
Tied up in knots: Untangling substrate recognition by the SPOUT methyltransferases.J Biol Chem. 2022 Oct;298(10):102393. doi: 10.1016/j.jbc.2022.102393. Epub 2022 Aug 18. J Biol Chem. 2022. PMID: 35988649 Free PMC article. Review.
-
Transfer RNA methyltransferases with a SpoU-TrmD (SPOUT) fold and their modified nucleosides in tRNA.Biomolecules. 2017 Feb 28;7(1):23. doi: 10.3390/biom7010023. Biomolecules. 2017. PMID: 28264529 Free PMC article. Review.
Cited by
-
Reviving the RNA World: An Insight into the Appearance of RNA Methyltransferases.Front Genet. 2016 Jun 6;7:99. doi: 10.3389/fgene.2016.00099. eCollection 2016. Front Genet. 2016. PMID: 27375676 Free PMC article. Review.
-
Methyltransferases: Functions and Applications.Chembiochem. 2022 Sep 16;23(18):e202200212. doi: 10.1002/cbic.202200212. Epub 2022 Jul 5. Chembiochem. 2022. PMID: 35691829 Free PMC article. Review.
-
Modifications and functional genomics of human transfer RNA.Cell Res. 2018 Apr;28(4):395-404. doi: 10.1038/s41422-018-0013-y. Epub 2018 Feb 20. Cell Res. 2018. PMID: 29463900 Free PMC article. Review.
-
Characterization of two homologous 2'-O-methyltransferases showing different specificities for their tRNA substrates.RNA. 2014 Aug;20(8):1257-71. doi: 10.1261/rna.044503.114. Epub 2014 Jun 20. RNA. 2014. PMID: 24951554 Free PMC article.
-
Structural and functional insights into tRNA binding and adenosine N1-methylation by an archaeal Trm10 homologue.Nucleic Acids Res. 2016 Jan 29;44(2):940-53. doi: 10.1093/nar/gkv1369. Epub 2015 Dec 15. Nucleic Acids Res. 2016. PMID: 26673726 Free PMC article.
References
-
- El Yacoubi B, Bailly M, de Crécy-Lagard V. Biosynthesis and function of posttranscriptional modifications of transfer RNAs. Annu. Rev. Genet. 2012;46:69–95. - PubMed
-
- Grosjean H. DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution. Austin, TX: Landes Bioscience; 2009.
-
- Phillips G, de Crécy-Lagard V. Biosynthesis and function of tRNA modifications in Archaea. Curr. Opin. Microbiol. 2011;14:335–341. - PubMed
-
- de Crécy-Lagard V, Marck C, Brochier-Armanet C, Grosjean H. Comparative RNomics and modomics in Mollicutes: prediction of gene function and evolutionary implications. IUBMB Life. 2007;59:634–658. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

