BRCA1-Dependent Translational Regulation in Breast Cancer Cells
- PMID: 23805307
- PMCID: PMC3689694
- DOI: 10.1371/journal.pone.0067313
BRCA1-Dependent Translational Regulation in Breast Cancer Cells
Abstract
BRCA1 (Breast Cancer 1) has been implicated in a number of cellular processes, including transcription regulation, DNA damage repair and protein ubiquitination. We previously demonstrated that BRCA1 interacts with PABP1 (Poly(A)-Binding Protein 1) and that BRCA1 modulates protein synthesis through this interaction. To identify the mRNAs that are translationally regulated by BRCA1, we used a microarray analysis of polysome-bound mRNAs in BRCA1-depleted and non-depleted MCF7 cells. Our findings show that BRCA1 modifies the translational efficiency of approximately 7% of the mRNAs expressed in these cells. Further analysis revealed that several processes contributing to cell surveillance such as cell cycle arrest, cell death, cellular growth and proliferation, DNA repair and gene expression, are largely enriched for the mRNAs whose translation is impacted by BRCA1. The BRCA1-dependent translation of these species of mRNAs therefore uncovers a novel mechanism through which BRCA1 exerts its onco-suppressive role. In addition, the BRCA1-dependent translation of mRNAs participating in unexpected functions such as cellular movement, nucleic acid metabolism or protein trafficking is indicative of novel functions for BRCA1. Finally, this study contributes to the identification of several markers associated with BRCA1 deficiency and to the discovery of new potential anti-neoplastic therapeutic targets.
Conflict of interest statement
Figures


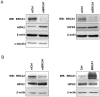
 ≤0.67, (↗)≥1.50, (↔) >0.67 and <1.50. The RRs are annotated with a sign and a number. The sign specifies the RR value: (−) ≤0.67, (+) ≥1.50. The number indicates how many mRNAs are deregulated.
≤0.67, (↗)≥1.50, (↔) >0.67 and <1.50. The RRs are annotated with a sign and a number. The sign specifies the RR value: (−) ≤0.67, (+) ≥1.50. The number indicates how many mRNAs are deregulated.  : mRNAs translationally deregulated through change in polysome mRNA abundance only;
: mRNAs translationally deregulated through change in polysome mRNA abundance only;  : mRNAs translationally deregulated through change in total mRNA abundance only;
: mRNAs translationally deregulated through change in total mRNA abundance only;  : mRNAs translationally deregulated through change in polysome abundance together with opposite changes in total mRNA C/Functional distribution of differentially translated known genes in BRCA1-depleted versus control MCF7 cells. Gene functions were established based on the annotation provided by the IPA database. The number of genes enriched in each function is shown in brackets.
: mRNAs translationally deregulated through change in polysome abundance together with opposite changes in total mRNA C/Functional distribution of differentially translated known genes in BRCA1-depleted versus control MCF7 cells. Gene functions were established based on the annotation provided by the IPA database. The number of genes enriched in each function is shown in brackets.
References
-
- Stratton MR, Rahman N (2008) The emerging landscape of breast cancer susceptibility. Nat Genet 40: 17–22. - PubMed
-
- Futreal PA, Liu Q, Shattuck-Eidens D, Cochran C, Harshman K, et al. (1994) BRCA1 mutations in primary breast and ovarian carcinomas. Science 266: 120–122. - PubMed
-
- Rio PG, Maurizis JC, Peffault de Latour M, Bignon YJ, Bernard-Gallon DJ (1999) Quantification of BRCA1 protein in sporadic breast carcinoma with or without loss of heterozygosity of the BRCA1 gene. Int J Cancer 80: 823–826. - PubMed
-
- Rakha EA, El-Sheikh SE, Kandil MA, El-Sayed ME, Green AR, et al. (2008) Expression of BRCA1 protein in breast cancer and its prognostic significance. Hum Pathol 39: 857–865. - PubMed
-
- Wilson CA, Ramos L, Villasenor MR, Anders KH, Press MF, et al. (1999) Localization of human BRCA1 and its loss in high-grade, non-inherited breast carcinomas. Nat Genet 21: 236–240. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

