Structure determination of membrane proteins in their native phospholipid bilayer environment by rotationally aligned solid-state NMR spectroscopy
- PMID: 23829871
- PMCID: PMC3778126
- DOI: 10.1021/ar400067z
Structure determination of membrane proteins in their native phospholipid bilayer environment by rotationally aligned solid-state NMR spectroscopy
Abstract
One of the most important topics in experimental structural biology is determining the structures of membrane proteins. These structures represent one-third of all of the information expressed from a genome, distinguished by their locations within the phospholipid bilayer of cells, organelles, or enveloped viruses. Their highly hydrophobic nature and insolubility in aqueous media means that they require an amphipathic environment. They have unique functions in transport, catalysis, channel formation, and signaling. Researchers are particularly interested in G-protein coupled receptors (GPCRs) because they modulate many biological processes, and about half of the approximately 800 of these proteins within the human genome are or can be turned into drug receptors that affect a wide range of diseases. Because of experimental difficulties, researchers have studied membrane proteins using a wide variety of artificial media that mimic membranes, such as mixed organic solvents or detergents. More sophisticated mimics include bilayer discs (bicelles) and the lipid cubic phase (LCP), but both of these contain a very large detergent component, which can disrupt the stability and function of membrane proteins. To have confidence in the resulting structures and their biological functions and to avoid disrupting these delicate proteins, the structures of membrane proteins should be determined in their native environment of liquid crystalline phospholipid bilayers under physiological conditions. This Account describes a recently developed general method for determining the structures of unmodified membrane proteins in phospholipid bilayers by solid-state NMR spectroscopy. Because it relies on the natural, rapid rotational diffusion of these proteins about the bilayer normal, this method is referred to as rotationally aligned (RA) solid-state NMR. This technique elaborates on oriented sample (OS) solid-state NMR, its complementary predecessor. These methods exploit the power of solid-state NMR, which enables researchers to obtain well-resolved spectra from "immobile" membrane proteins in phospholipid bilayers, to separate and measure frequencies that reflect orientations with respect to the bilayer normal, and to make complementary distance measurements. The determination of the structures of several membrane proteins, most prominently the chemokine receptor CXCR1, a 350-residue GPCR, has demonstrated this approach.
Figures

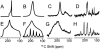



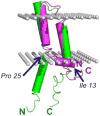

References
-
- Dickerson RE, Reddy J, Pinkerton M, Steinrauf LK. A 6 angstrom model of triclinic lysozyme. Nature. 1962;196:1178. - PubMed
-
- Kendrew JC, Bodo G, Dintzis HM, Parrish RG, Wyckoff HW, Philips DC. A three-dimensiona model of the myoglobin molecule obtained by X-ray analysis. Nature. 1958;181:666. - PubMed
-
- Perutz MF, Rossman MG, Cullis AF, Muirhead H, Georg W. Structure of hemoglobin: A three-dimensional Fourier synthesis at 5.5 A resolution, obtaine by X-ray analyss. Nature. 1960;185:416–422. - PubMed
-
- Saunders M, Wishnia A, Kirkwood JG. The Nuclear Magnetic Resonance Spectrum of Ribonuclease1. Journal of the American Chemical Society. 1957;79:3289–3290.
-
- Anfinsen CB. Principles that govern the folding of protein chains. Science. 1973;181:223–30. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

