The FACT histone chaperone guides histone H4 into its nucleosomal conformation in Saccharomyces cerevisiae
- PMID: 23833181
- PMCID: PMC3761294
- DOI: 10.1534/genetics.113.153080
The FACT histone chaperone guides histone H4 into its nucleosomal conformation in Saccharomyces cerevisiae
Abstract
The pob3-Q308K mutation alters the small subunit of the Saccharomyces cerevisiae histone/nucleosome chaperone Facilitates Chromatin Transactions (FACT), causing defects in both transcription and DNA replication. We describe histone mutations that suppress some of these defects, providing new insight into the mechanism of FACT activity in vivo. FACT is primarily known for its ability to promote reorganization of nucleosomes into a more open form, but neither the pob3-Q308K mutation nor the compensating histone mutations affect this activity. Instead, purified mutant FACT complexes fail to release from nucleosomes efficiently, and the histone mutations correct this flaw. We confirm that pob3-T252E also suppresses pob3-Q308K and show that combining two suppressor mutations can be detrimental, further demonstrating the importance of balance between association and dissociation for efficient FACT:nucleosome interactions. To explain our results, we propose that histone H4 can adopt multiple conformations, most of which are incompatible with nucleosome assembly. FACT guides H4 to adopt appropriate conformations, and this activity can be enhanced or diminished by mutations in Pob3 or histones. FACT can therefore destabilize nucleosomes by favoring the reorganized state, but it can also promote assembly by tethering histones and DNA together and maintaining them in conformations that promote canonical nucleosome formation.
Keywords: FACT suppressor; histone mutations; histone rearrangement; nucleosome reorganization.
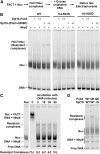
Figures

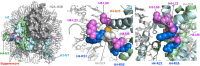
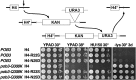
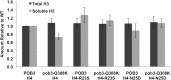
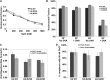


Similar articles
-
The Abundant Histone Chaperones Spt6 and FACT Collaborate to Assemble, Inspect, and Maintain Chromatin Structure in Saccharomyces cerevisiae.Genetics. 2015 Nov;201(3):1031-45. doi: 10.1534/genetics.115.180794. Epub 2015 Sep 28. Genetics. 2015. PMID: 26416482 Free PMC article.
-
Insight into the mechanism of nucleosome reorganization from histone mutants that suppress defects in the FACT histone chaperone.Genetics. 2011 Aug;188(4):835-46. doi: 10.1534/genetics.111.128769. Epub 2011 May 30. Genetics. 2011. PMID: 21625001 Free PMC article.
-
The Histone Chaperone FACT Contributes to DNA Replication-Coupled Nucleosome Assembly.Cell Rep. 2016 Feb 9;14(5):1128-1141. doi: 10.1016/j.celrep.2015.12.096. Epub 2016 Jan 21. Cell Rep. 2016. PMID: 26804921
-
[Structure and function of histone chaperone FACT].Mol Biol (Mosk). 2015 Nov-Dec;49(6):891-904. doi: 10.7868/S0026898415060026. Mol Biol (Mosk). 2015. PMID: 26710768 Review. Russian.
-
Replication-Coupled Nucleosome Assembly in the Passage of Epigenetic Information and Cell Identity.Trends Biochem Sci. 2018 Feb;43(2):136-148. doi: 10.1016/j.tibs.2017.12.003. Epub 2017 Dec 29. Trends Biochem Sci. 2018. PMID: 29292063 Free PMC article. Review.
Cited by
-
The Abundant Histone Chaperones Spt6 and FACT Collaborate to Assemble, Inspect, and Maintain Chromatin Structure in Saccharomyces cerevisiae.Genetics. 2015 Nov;201(3):1031-45. doi: 10.1534/genetics.115.180794. Epub 2015 Sep 28. Genetics. 2015. PMID: 26416482 Free PMC article.
-
The CENP-T/-W complex is a binding partner of the histone chaperone FACT.Genes Dev. 2016 Jun 1;30(11):1313-26. doi: 10.1101/gad.275073.115. Epub 2016 Jun 9. Genes Dev. 2016. PMID: 27284163 Free PMC article.
-
Establishment and Maintenance of Chromatin Architecture Are Promoted Independently of Transcription by the Histone Chaperone FACT and H3-K56 Acetylation in Saccharomyces cerevisiae.Genetics. 2019 Mar;211(3):877-892. doi: 10.1534/genetics.118.301853. Epub 2019 Jan 24. Genetics. 2019. PMID: 30679261 Free PMC article.
-
Functional roles of the DNA-binding HMGB domain in the histone chaperone FACT in nucleosome reorganization.J Biol Chem. 2018 Apr 20;293(16):6121-6133. doi: 10.1074/jbc.RA117.000199. Epub 2018 Mar 7. J Biol Chem. 2018. PMID: 29514976 Free PMC article.
-
Pluripotency state transition of embryonic stem cells requires the turnover of histone chaperone FACT on chromatin.iScience. 2023 Nov 28;27(1):108537. doi: 10.1016/j.isci.2023.108537. eCollection 2024 Jan 19. iScience. 2023. PMID: 38213626 Free PMC article.
References
-
- Ahn S. H., Cheung W. L., Hsu J. Y., Diaz R. L., Smith M. M., et al. , 2005. Sterile 20 kinase phosphorylates histone H2B at serine 10 during hydrogen peroxide-induced apoptosis in S. cerevisiae. Cell 120: 25–36. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

