Membrane elastic properties and cell function
- PMID: 23844071
- PMCID: PMC3701085
- DOI: 10.1371/journal.pone.0067708
Membrane elastic properties and cell function
Abstract
Recent studies indicate that the cell membrane, interacting with its attached cytoskeleton, is an important regulator of cell function, exerting and responding to forces. We investigate this relationship by looking for connections between cell membrane elastic properties, especially surface tension and bending modulus, and cell function. Those properties are measured by pulling tethers from the cell membrane with optical tweezers. Their values are determined for all major cell types of the central nervous system, as well as for macrophage. Astrocytes and glioblastoma cells, which are considerably more dynamic than neurons, have substantially larger surface tensions. Resting microglia, which continually scan their environment through motility and protrusions, have the highest elastic constants, with values similar to those for resting macrophage. For both microglia and macrophage, we find a sharp softening of bending modulus between their resting and activated forms, which is very advantageous for their acquisition of phagocytic functions upon activation. We also determine the elastic constants of pure cell membrane, with no attached cytoskeleton. For all cell types, the presence of F-actin within tethers, contrary to conventional wisdom, is confirmed. Our findings suggest the existence of a close connection between membrane elastic constants and cell function.
Conflict of interest statement
Figures
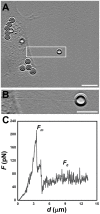
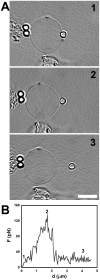
 , extracted from both Neuron CX and Neuron GE in regions 1, 2 and 3. (H) SEM image of a typical tether extracted from Neuron GE. Scale bar is 1 µm. (I) Mean values of the tether radius, R, extracted from both Neuron CX and Neuron GE in regions 1, 2 and 3. Standard errors were used as error bars in (G) and (H). At least 20 different experiments were performed for each situation in (G) and (H). (J)–(L) Images of tethers extracted from Neuron GE stained for F-actin with phaloidin-FITC, in green (J), β-tubulin III, in red (K) and the merge of both images (L). Scale bar is 5 µm.
, extracted from both Neuron CX and Neuron GE in regions 1, 2 and 3. (H) SEM image of a typical tether extracted from Neuron GE. Scale bar is 1 µm. (I) Mean values of the tether radius, R, extracted from both Neuron CX and Neuron GE in regions 1, 2 and 3. Standard errors were used as error bars in (G) and (H). At least 20 different experiments were performed for each situation in (G) and (H). (J)–(L) Images of tethers extracted from Neuron GE stained for F-actin with phaloidin-FITC, in green (J), β-tubulin III, in red (K) and the merge of both images (L). Scale bar is 5 µm.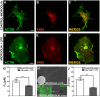
 , extracted from microglial cells. (H) SEM image of a typical tether extracted from microglia+LPS. Scale bar is 1 µm. (I) Image of tether extracted from control microglia stained for F-actin with phaloidin-FITC, in green. Scale bar is 5 µm. (J) Mean values of the tether radius, R, extracted from both microglial cell conditions. Standard errors were used as error bars in (G) and (J). At least 20 different experiments were performed for each situation in (G) and (J) (*** means p<0.0001 in t-test statistics).
, extracted from microglial cells. (H) SEM image of a typical tether extracted from microglia+LPS. Scale bar is 1 µm. (I) Image of tether extracted from control microglia stained for F-actin with phaloidin-FITC, in green. Scale bar is 5 µm. (J) Mean values of the tether radius, R, extracted from both microglial cell conditions. Standard errors were used as error bars in (G) and (J). At least 20 different experiments were performed for each situation in (G) and (J) (*** means p<0.0001 in t-test statistics).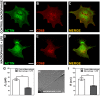
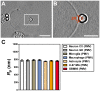
 , extracted from macrophage cells in both conditions. (H) SEM image of a typical tether extracted from macrophage+LPS cells. Scale bar is 1 µm. (I) Mean values of the tether radius, R, extracted from macrophage cells in both conditions. Standard errors were used as error bars in (G) and (I). At least 20 different experiments were performed for each situation in (G) and (I). (*** means p<0.0001 in t-test statistics).
, extracted from macrophage cells in both conditions. (H) SEM image of a typical tether extracted from macrophage+LPS cells. Scale bar is 1 µm. (I) Mean values of the tether radius, R, extracted from macrophage cells in both conditions. Standard errors were used as error bars in (G) and (I). At least 20 different experiments were performed for each situation in (G) and (I). (*** means p<0.0001 in t-test statistics).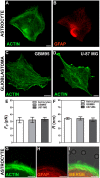
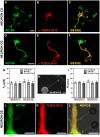
 , extracted from astrocytes and glioblastomas cells. (F) Mean values of the tether radius,R, extracted from astrocytes and glioblastomas cells. Standard errors were used as error bars in (E) and (F). At least 20 different experiments were performed for each situation in (E) and (F). (G-I) Images of tethers extracted from astrocytes, stained for F-actin with phaloidin-FITC, in green (G) and GFAP, in red (H). (G) and (H) merged in (I). Scale bars for G-I are all 5 µm.
, extracted from astrocytes and glioblastomas cells. (F) Mean values of the tether radius,R, extracted from astrocytes and glioblastomas cells. Standard errors were used as error bars in (E) and (F). At least 20 different experiments were performed for each situation in (E) and (F). (G-I) Images of tethers extracted from astrocytes, stained for F-actin with phaloidin-FITC, in green (G) and GFAP, in red (H). (G) and (H) merged in (I). Scale bars for G-I are all 5 µm.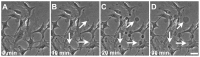
References
-
- Sheetz MP (2001) Cell control by membrane-cytoskeleton adhesion. Nat Rev Mol Cell Biol 2: 392–396. - PubMed
-
- Janmey PA, McCulloch CA (2007) Cell mechanics: integrating cell responses to mechanical stimuli. Annu Rev Biomed Eng 9: 1–34. - PubMed
-
- Flannagan RS, Jaumouillé V, Grinstein S (2012) The cell biology of phagocytosis. Ann Rev Pathol 7: 61–98. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

