Development of a magnetic electrochemical bar code array for point mutation detection in the H5N1 neuraminidase gene
- PMID: 23860384
- PMCID: PMC3738958
- DOI: 10.3390/v5071719
Development of a magnetic electrochemical bar code array for point mutation detection in the H5N1 neuraminidase gene
Abstract
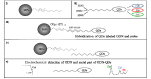
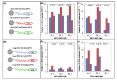
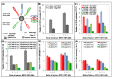
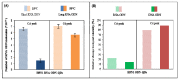
Since its first official detection in the Guangdong province of China in 1996, the highly pathogenic avian influenza virus of H5N1 subtype (HPAI H5N1) has reportedly been the cause of outbreaks in birds in more than 60 countries, 24 of which were European. The main issue is still to develop effective antiviral drugs. In this case, single point mutation in the neuraminidase gene, which causes resistance to antiviral drug and is, therefore, subjected to many studies including ours, was observed. In this study, we developed magnetic electrochemical bar code array for detection of single point mutations (mismatches in up to four nucleotides) in H5N1 neuraminidase gene. Paramagnetic particles Dynabeads® with covalently bound oligo (dT)₂₅ were used as a tool for isolation of complementary H5N1 chains (H5N1 Zhejin, China and Aichi). For detection of H5N1 chains, oligonucleotide chains of lengths of 12 (+5 adenine) or 28 (+5 adenine) bp labeled with quantum dots (CdS, ZnS and/or PbS) were used. Individual probes hybridized to target molecules specifically with efficiency higher than 60%. The obtained signals identified mutations present in the sequence. Suggested experimental procedure allows obtaining further information from the redox signals of nucleic acids. Moreover, the used biosensor exhibits sequence specificity and low limits of detection of subnanogram quantities of target nucleic acids.
Figures

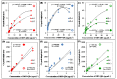
Similar articles
-
The significance of naturally occurring neuraminidase quasispecies of H5N1 avian influenza virus on resistance to oseltamivir: a point of concern.J Gen Virol. 2016 Jun;97(6):1311-1323. doi: 10.1099/jgv.0.000444. Epub 2016 Mar 2. J Gen Virol. 2016. PMID: 26935590 Free PMC article.
-
Neuraminidase inhibitor-resistant recombinant A/Vietnam/1203/04 (H5N1) influenza viruses retain their replication efficiency and pathogenicity in vitro and in vivo.J Virol. 2007 Nov;81(22):12418-26. doi: 10.1128/JVI.01067-07. Epub 2007 Sep 12. J Virol. 2007. PMID: 17855542 Free PMC article.
-
Therapeutic efficacy of peramivir against H5N1 highly pathogenic avian influenza viruses harboring the neuraminidase H275Y mutation.Antiviral Res. 2017 Mar;139:41-48. doi: 10.1016/j.antiviral.2016.12.011. Epub 2016 Dec 22. Antiviral Res. 2017. PMID: 28012921
-
Rapid quantitation of neuraminidase inhibitor drug resistance in influenza virus quasispecies.Antivir Ther. 2008;13(6):809-20. Antivir Ther. 2008. PMID: 18839782
-
Molecular mechanisms underlying oseltamivir resistance mediated by an I117V substitution in the neuraminidase of subtype H5N1 avian influenza A viruses.J Infect Dis. 2013 Jan 1;207(1):89-97. doi: 10.1093/infdis/jis633. Epub 2012 Oct 10. J Infect Dis. 2013. PMID: 23053629 Free PMC article.
Cited by
-
Biosensors for Point Mutation Detection.Front Bioeng Biotechnol. 2021 Dec 15;9:797831. doi: 10.3389/fbioe.2021.797831. eCollection 2021. Front Bioeng Biotechnol. 2021. PMID: 34976987 Free PMC article. No abstract available.
-
Nanomaterial-based biosensors for detection of pathogenic virus.Trends Analyt Chem. 2017 Dec;97:445-457. doi: 10.1016/j.trac.2017.10.005. Epub 2017 Oct 13. Trends Analyt Chem. 2017. PMID: 32287543 Free PMC article. Review.
References
-
- Wei K.F., Chen Y.F., Chen J., Wu L.J., Xie D.X. Evolution and adaptation of hemagglutinin gene of human H5N1 influenza virus. Virus Genes. 2012;44:450–458. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

