Nuclear dynamics in a fungal chimera
- PMID: 23861490
- PMCID: PMC3740868
- DOI: 10.1073/pnas.1220842110
Nuclear dynamics in a fungal chimera
Abstract
A fungal colony is a syncytium composed of a branched and interconnected network of cells. Chimerism endows colonies with increased virulence and ability to exploit nutritionally complex substrates. Moreover, chimera formation may be a driver for diversification at the species level by allowing lateral gene transfer between strains that are too distantly related to hybridize sexually. However, the processes by which genomic diversity develops and is maintained within a single colony are little understood. In particular, both theory and experiments show that genetically diverse colonies may be unstable and spontaneously segregate into genetically homogenous sectors. By directly measuring patterns of nuclear movement in the model ascomycete fungus Neurospora crassa, we show that genetic diversity is maintained by complex mixing flows of nuclei at all length scales within the hyphal network. Mathematical modeling and experiments in a morphological mutant reveal some of the exquisite hydraulic engineering necessary to create the mixing flows. In addition to illuminating multinucleate and multigenomic lifestyles, the adaptation of a hyphal network for mixing nuclear material provides a previously unexamined organizing principle for understanding morphological diversity in the more-than-a-million species of filamentous fungi.
Keywords: biological networks; heterokaryon; hydrodynamics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
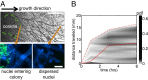

 , velocity pdf under osmotic gradient (green) is the same as for normal growth (blue). (Scale bars, 50 μm.)
, velocity pdf under osmotic gradient (green) is the same as for normal growth (blue). (Scale bars, 50 μm.)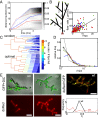
 . (C) The probability
. (C) The probability  of sibling nuclei being sent to different tips was optimized by Monte Carlo simulations (
of sibling nuclei being sent to different tips was optimized by Monte Carlo simulations ( from 0.37 in a random branching network (Upper) to a value close to 0.46 (Lower). Branches are color coded by their flow rates. (D) For real colonies the distribution of branches at each stage of the hierarchy (blue, 3-cm mycelium; green, 4 cm; red, 5 cm) is close to optimal (solid black curve and crosses) rather than random branching (dashed black curve). (E) Despite having close to optimal branching, a so chimera becomes unmixed with growth. Conidial chains of a his-3::hH1-gfp; Pccg1-DsRed so + his-3::hH1-gfp; so heterokaryon tend to contain only hH1-GFP so nuclei (Left) or hH1-GFP DsRed so nuclei (Center); compare a heterokaryotic wild-type conidial chain in which hH1-DsRed and hH1-GFP nuclei are evenly mixed (Upper Right). (Scale bars, 20 μm.) Graph showing narrow spread of
from 0.37 in a random branching network (Upper) to a value close to 0.46 (Lower). Branches are color coded by their flow rates. (D) For real colonies the distribution of branches at each stage of the hierarchy (blue, 3-cm mycelium; green, 4 cm; red, 5 cm) is close to optimal (solid black curve and crosses) rather than random branching (dashed black curve). (E) Despite having close to optimal branching, a so chimera becomes unmixed with growth. Conidial chains of a his-3::hH1-gfp; Pccg1-DsRed so + his-3::hH1-gfp; so heterokaryon tend to contain only hH1-GFP so nuclei (Left) or hH1-GFP DsRed so nuclei (Center); compare a heterokaryotic wild-type conidial chain in which hH1-DsRed and hH1-GFP nuclei are evenly mixed (Upper Right). (Scale bars, 20 μm.) Graph showing narrow spread of  between wild-type conidial chains (black line) indicates more mixing of nucleotypes than in so (dashed red line).
between wild-type conidial chains (black line) indicates more mixing of nucleotypes than in so (dashed red line).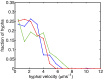
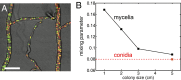
 , independent of colony size (blue, 3-cm mycelium; green, 4 cm; red, 5 cm). We use this result to estimate the variance in travel times for sibling nuclei traveling from the colony interior to a growing hyphal tip (main text).
, independent of colony size (blue, 3-cm mycelium; green, 4 cm; red, 5 cm). We use this result to estimate the variance in travel times for sibling nuclei traveling from the colony interior to a growing hyphal tip (main text).References
-
- Booy G, Hendirks R, Dmulders M, Van Groenendael J, Vosman B. Genetic diversity and the survival of populations. Plant Biol. 2000;2:379–395.
-
- Crutsinger GM, et al. Plant genotypic diversity predicts community structure and governs an ecosystem process. Science. 2006;313(5789):966–968. - PubMed
-
- Gray W, Alexopoulos C. Biology of the Myxomycetes. New York: Ronald Press; 1968.
-
- Maheshwari R. Nuclear behavior in fungal hyphae. FEMS Microbiol Lett. 2005;249(1):7–14. - PubMed
-
- Sidhu G. Genetics of Giberella fujikuroi. iii. Significance of heterokaryosis in naturally occurring corn isolates. Can J Bot. 1983;61:3320–3325.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

