The template specificity of bacteriophage Phi6 RNA polymerase
- PMID: 23864621
- PMCID: PMC3754000
- DOI: 10.1128/JVI.01467-13
The template specificity of bacteriophage Phi6 RNA polymerase
Abstract
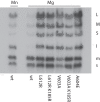
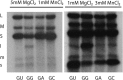
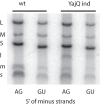
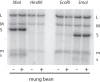
Bacteriophage Φ6 contains three double-stranded RNA (dsRNA) genomic segments, L, M, and S. The RNA is located inside a core particle composed of multiple copies of a major structural protein, an RNA-dependent RNA polymerase, a hexameric NTPase, and an auxiliary protein. The virion RNA polymerase in the core particle transcribes segments M and S in vitro. Segment L is transcribed poorly because its transcript starts with GU instead of GG found on segments S and M. Transcription in vivo is modified by the binding of host protein YajQ to the outside the core particle so that segment L is transcribed well. This mechanism is the determinant of the temporal control of gene expression in Φ6. Mutants of Φ6 have been isolated that are independent of YajQ for transcription of segment L. The mutations are found in the gene of the viral polymerase or the major capsid protein or both. These mutants are capable of transcribing segment L with the GU start or GA or GC. The same is found to be true when YajQ is added to wild-type particles. Minus-strand synthesis has restrictions that are different from that of plus-strand synthesis, and YajQ or mutations to independence do not modify minus-strand synthesis behavior. Purified polymerase P2 is able to transcribe dsRNA, but transcription behavior of segment L by both wild-type and mutant polymerases is different from that seen in capsid structures. Adding YajQ to purified polymerase does not change its transcription specificity.
Figures



Similar articles
-
Structural Studies of Bacteriophage Φ6 and Its Transformations during Its Life Cycle.Viruses. 2023 Dec 11;15(12):2404. doi: 10.3390/v15122404. Viruses. 2023. PMID: 38140645 Free PMC article. Review.
-
The role of host protein YajQ in the temporal control of transcription in bacteriophage Phi6.Proc Natl Acad Sci U S A. 2008 Oct 14;105(41):15956-60. doi: 10.1073/pnas.0807489105. Epub 2008 Oct 3. Proc Natl Acad Sci U S A. 2008. PMID: 18836083 Free PMC article.
-
Protein P7 of phage phi6 RNA polymerase complex, acquiring of RNA packaging activity by in vitro assembly of the purified protein onto deficient particles.J Mol Biol. 1997 Mar 14;266(5):891-900. doi: 10.1006/jmbi.1996.0817. J Mol Biol. 1997. PMID: 9086268
-
Comparison of polymerase subunits from double-stranded RNA bacteriophages.J Virol. 2001 Nov;75(22):11088-95. doi: 10.1128/JVI.75.22.11088-11095.2001. J Virol. 2001. PMID: 11602748 Free PMC article.
-
Precise packaging of the three genomic segments of the double-stranded-RNA bacteriophage phi6.Microbiol Mol Biol Rev. 1999 Mar;63(1):149-60. doi: 10.1128/MMBR.63.1.149-160.1999. Microbiol Mol Biol Rev. 1999. PMID: 10066834 Free PMC article. Review.
Cited by
-
Discovery and Classification of the φ6 Bacteriophage: An Historical Review.Viruses. 2023 May 31;15(6):1308. doi: 10.3390/v15061308. Viruses. 2023. PMID: 37376608 Free PMC article. Review.
-
Cystoviral RNA-directed RNA polymerases: Regulation of RNA synthesis on multiple time and length scales.Virus Res. 2017 Apr 15;234:135-152. doi: 10.1016/j.virusres.2017.01.006. Epub 2017 Jan 16. Virus Res. 2017. PMID: 28104452 Free PMC article. Review.
-
RNA Packaging in the Cystovirus Bacteriophages: Dynamic Interactions during Capsid Maturation.Int J Mol Sci. 2022 Feb 28;23(5):2677. doi: 10.3390/ijms23052677. Int J Mol Sci. 2022. PMID: 35269819 Free PMC article. Review.
-
Cystoviral polymerase complex protein P7 uses its acidic C-terminal tail to regulate the RNA-directed RNA polymerase P2.J Mol Biol. 2014 Jul 15;426(14):2580-93. doi: 10.1016/j.jmb.2014.04.028. Epub 2014 May 9. J Mol Biol. 2014. PMID: 24813120 Free PMC article.
-
Structural Studies of Bacteriophage Φ6 and Its Transformations during Its Life Cycle.Viruses. 2023 Dec 11;15(12):2404. doi: 10.3390/v15122404. Viruses. 2023. PMID: 38140645 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

