Subnuclear partitioning of rRNA genes between the nucleolus and nucleoplasm reflects alternative epiallelic states
- PMID: 23873938
- PMCID: PMC3731543
- DOI: 10.1101/gad.221648.113
Subnuclear partitioning of rRNA genes between the nucleolus and nucleoplasm reflects alternative epiallelic states
Abstract
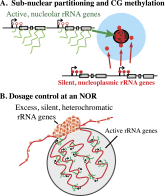
Eukaryotes can have thousands of 45S ribosomal RNA (rRNA) genes, many of which are silenced during development. Using fluorescence-activated sorting techniques, we show that active rRNA genes in Arabidopsis thaliana are present within sorted nucleoli, whereas silenced rRNA genes are excluded. DNA methyltransferase (met1), histone deacetylase (hda6), or chromatin assembly (caf1) mutants that disrupt silencing abrogate this nucleoplasmic-nucleolar partitioning. Bisulfite sequencing data indicate that active nucleolar rRNA genes are nearly completely demethylated at promoter CGs, whereas silenced genes are nearly fully methylated. Collectively, the data reveal that rRNA genes occupy distinct but changeable nuclear territories according to their epigenetic state.
Keywords: DNA methylation; RNA polymerase I; chromatin assembly; gene silencing; histone deacetylation; ribosomal RNA gene; transcription.
Figures



References
-
- Barneche F, Steinmetz F, Echeverria M 2000. Fibrillarin genes encode both a conserved nucleolar protein and a novel small nucleolar RNA involved in ribosomal RNA methylation in Arabidopsis thaliana. J Biol Chem 275: 27212–27220 - PubMed
-
- Caperta AD, Neves N, Morais-Cecilio L, Malho R, Viegas W 2002. Genome restructuring in rye affects the expression, organization and disposition of homologous rDNA loci. J Cell Sci 115: 2839–2846 - PubMed
-
- Copenhaver GP, Pikaard CS 1996. RFLP and physical mapping with an rDNA-specific endonuclease reveals that nucleolus organizer regions of Arabidopsis thaliana adjoin the telomeres on chromosomes 2 and 4. Plant J 9: 259–272 - PubMed
-
- Copenhaver GP, Doelling JH, Gens S, Pikaard CS 1995. Use of RFLPs larger than 100 kbp to map the position and internal organization of the nucleolus organizer region on chromosome 2 in Arabidopsis thaliana. Plant J 7: 273–286 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
