Tyrosine Y189 in the substrate domain of the adhesion docking protein NEDD9 is conserved with p130Cas Y253 and regulates NEDD9-mediated migration and focal adhesion dynamics
- PMID: 23874939
- PMCID: PMC3706375
- DOI: 10.1371/journal.pone.0069304
Tyrosine Y189 in the substrate domain of the adhesion docking protein NEDD9 is conserved with p130Cas Y253 and regulates NEDD9-mediated migration and focal adhesion dynamics
Abstract
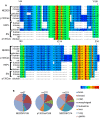
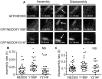
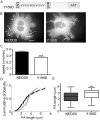
The focal adhesion docking protein NEDD9/HEF1/Cas-L regulates cell migration and cancer invasion. NEDD9 is a member of the Cas family of proteins that share conserved overall protein-protein interaction domain structure, including a substrate domain that is characterized by extensive tyrosine (Y) phosphorylation. Previous studies have suggested that phosphorylation of Y253 in the substrate domain of the Cas family protein p130Cas is specifically required for p130Cas function in cell migration. While it is clear that tyrosine phosphorylation of the NEDD9 substrate domain is similarly required for the regulation of cell motility, whether individual NEDD9 tyrosine residues have discrete function in regulating motility has not previously been reported. In the present study we have used a global sequence alignment of Cas family proteins to identify a putative NEDD9 equivalent of p130Cas Y253. We find that NEDD9 Y189 aligns with p130Cas Y253 and that it is conserved among NEDD9 vertebrate orthologues. Expression of NEDD9 in which Y189 is mutated to phenylalanine results in increased rates of cell migration and is correlated with increased disassembly of GFP.NEDD9 focal adhesions. Conversely, mutation to Y189D significantly inhibits cell migration. Our previous data has suggested that NEDD9 stabilizes focal adhesions and the present data therefore suggests that phosphorylation of Y189 NEDD9 is required for this function. These findings indicate that the individual tyrosine residues of the NEDD9 substrate domain may serve discrete functional roles. Given the important role of this protein in promoting cancer invasion, greater understanding of the function of the individual tyrosine residues is important for the future design of approaches to target NEDD9 to arrest cancer cell invasion.
Conflict of interest statement
Figures


Similar articles
-
The focal adhesion targeting domain of p130Cas confers a mechanosensing function.J Cell Sci. 2017 Apr 1;130(7):1263-1273. doi: 10.1242/jcs.192930. Epub 2017 Feb 21. J Cell Sci. 2017. PMID: 28223315
-
P130Cas Src-binding and substrate domains have distinct roles in sustaining focal adhesion disassembly and promoting cell migration.PLoS One. 2010 Oct 18;5(10):e13412. doi: 10.1371/journal.pone.0013412. PLoS One. 2010. PMID: 20976150 Free PMC article.
-
A novel Cas family member, HEPL, regulates FAK and cell spreading.Mol Biol Cell. 2008 Apr;19(4):1627-36. doi: 10.1091/mbc.e07-09-0953. Epub 2008 Feb 6. Mol Biol Cell. 2008. PMID: 18256281 Free PMC article.
-
Molecular basis for HEF1/NEDD9/Cas-L action as a multifunctional co-ordinator of invasion, apoptosis and cell cycle.Cell Biochem Biophys. 2007;48(1):54-72. doi: 10.1007/s12013-007-0036-3. Cell Biochem Biophys. 2007. PMID: 17703068 Free PMC article. Review.
-
CAS proteins in normal and pathological cell growth control.Cell Mol Life Sci. 2010 Apr;67(7):1025-48. doi: 10.1007/s00018-009-0213-1. Epub 2009 Nov 25. Cell Mol Life Sci. 2010. PMID: 19937461 Free PMC article. Review.
Cited by
-
CAS proteins in health and disease: an update.IUBMB Life. 2014 Jun;66(6):387-95. doi: 10.1002/iub.1282. Epub 2014 Jun 24. IUBMB Life. 2014. PMID: 24962474 Free PMC article. Review.
-
Macrophages downregulate NEDD9 to counteract S. Typhimurium- mediated FAK-AKT activation and lysosome inhibition.Cell Death Dis. 2025 Jun 12;16(1):445. doi: 10.1038/s41419-025-07634-9. Cell Death Dis. 2025. PMID: 40506423 Free PMC article.
-
The scaffold protein CasL restrains membrane blebbing and promotes T cell migration.J Cell Sci. 2025 Jul 15;138(14):jcs263792. doi: 10.1242/jcs.263792. Epub 2025 Jul 18. J Cell Sci. 2025. PMID: 40576420
-
SOX9 is a dose-dependent metastatic fate determinant in melanoma.J Exp Clin Cancer Res. 2019 Jan 14;38(1):17. doi: 10.1186/s13046-018-0998-6. J Exp Clin Cancer Res. 2019. PMID: 30642390 Free PMC article.
-
NEDD9 stimulated MMP9 secretion is required for invadopodia formation in oral squamous cell carcinoma.Oncotarget. 2018 May 22;9(39):25503-25516. doi: 10.18632/oncotarget.25347. eCollection 2018 May 22. Oncotarget. 2018. PMID: 29876004 Free PMC article.
References
-
- Sanz-Moreno V, Gadea G, Ahn J, Paterson H, Marra P, et al. (2008) Rac activation and inactivation control plasticity of tumor cell movement. Cell 135: 510–523. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

