Three-dimensional reconstruction of icosahedral particles from single micrographs in real time at the microscope
- PMID: 23891839
- PMCID: PMC3831522
- DOI: 10.1016/j.jsb.2013.07.007
Three-dimensional reconstruction of icosahedral particles from single micrographs in real time at the microscope
Abstract
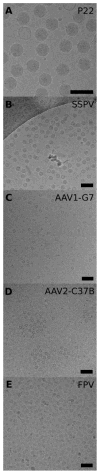
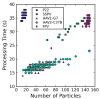
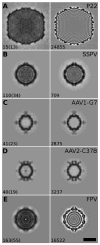
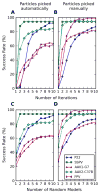
Single particle analysis is a valuable tool in cryo-electron microscopy for determining the structure of biological complexes. However, the conformational state and the preparation of the sample are factors that play a critical role in the ultimate attainable resolution. In some cases extensive analysis at the microscope of a sample under different conditions is required to derive the optimal acquisition conditions. Currently this analysis is limited to raw micrographs, thus conveying only limited information on the structure of the complex. We are developing a computing system that generates a three-dimensional reconstruction from a single micrograph acquired under cryogenic and low dose conditions, and containing particles with icosahedral symmetry. The system provides the microscopist with immediate structural information from a sample while it is in the microscope and during the preliminary acquisition stage. The system is designed to run without user intervention on a multi-processor computing resource and integrates all the processing steps required for the analysis. Tests performed on experimental data sets show that the probability of obtaining a reliable reconstruction from one micrograph is primarily determined by the quality of the sample, with success rates close to 100% when sample conditions are optimal, and decreasing to about 60% when conditions are sub-optimal. The time required to generate a reconstruction depends significantly on the diameter of the particles, and in most instances takes about 1min. The proposed approach can provide valuable three-dimensional information, albeit at low resolution, on conformational states, epitope binding, and stoichiometry of icosahedral multi-protein complexes.
Keywords: 3D reconstruction; Automation; Cryo-electron microscopy; Icosahedral particles; Image processing; Single particle analysis.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures




Similar articles
-
Single particle analysis integrated with microscopy: a high-throughput approach for reconstructing icosahedral particles.J Struct Biol. 2014 Apr;186(1):8-18. doi: 10.1016/j.jsb.2014.02.016. Epub 2014 Mar 5. J Struct Biol. 2014. PMID: 24613762 Free PMC article.
-
Structure of HIV-1 capsid assemblies by cryo-electron microscopy and iterative helical real-space reconstruction.J Vis Exp. 2011 Aug 9;(54):3041. doi: 10.3791/3041. J Vis Exp. 2011. PMID: 21860371 Free PMC article.
-
Single-particle reconstruction statistics: a diagnostic tool in solving biomolecular structures by cryo-EM.Acta Crystallogr F Struct Biol Commun. 2019 Jan 1;75(Pt 1):33-44. doi: 10.1107/S2053230X18017636. Epub 2019 Jan 1. Acta Crystallogr F Struct Biol Commun. 2019. PMID: 30605123 Free PMC article. Review.
-
Solution x-ray scattering-based estimation of electron cryomicroscopy imaging parameters for reconstruction of virus particles.Biophys J. 1999 Apr;76(4):2249-61. doi: 10.1016/S0006-3495(99)77381-9. Biophys J. 1999. PMID: 10096920 Free PMC article.
-
Three-dimensional electron cryo-microscopy as a powerful structural tool in molecular medicine.J Mol Med (Berl). 2000;78(4):191-202. doi: 10.1007/s001090000101. J Mol Med (Berl). 2000. PMID: 10933581 Review.
Cited by
-
Validation of 3D EM Reconstructions: The Phantom in the Noise.AIMS Biophys. 2015;2(1):21-35. doi: 10.3934/biophy.2015.1.21. AIMS Biophys. 2015. PMID: 26005714 Free PMC article.
-
Single particle analysis integrated with microscopy: a high-throughput approach for reconstructing icosahedral particles.J Struct Biol. 2014 Apr;186(1):8-18. doi: 10.1016/j.jsb.2014.02.016. Epub 2014 Mar 5. J Struct Biol. 2014. PMID: 24613762 Free PMC article.
References
-
- Adiga U, Baxter WT, Hall RJ, Rockel B, Rath BK, Frank J, Glaeser R. Particle picking by segmentation: A comparative study with SPIDER-based manual particle picking. J Struct Biol. 2005;152:211–220. - PubMed
-
- Baker TS, Cheng RH. A model-based approach for determining orientations of biological macromolecules imaged by cryoelectron microscopy. J Struct Biol. 1996;116:120–130. - PubMed
-
- Boier Martin IM, Marinescu DC, Lynch RE, Baker TS. Identification of spherical virus particles in digitized images of entire electron micrographs. J Struct Biol. 1997;120:146–157. - PubMed
-
- Frank J. Three-Dimensional Electron Microscopy of Macromolecular Assemblies. Oxford U. Press; 2006.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

