The scavenger receptor SCARF1 mediates the clearance of apoptotic cells and prevents autoimmunity
- PMID: 23892722
- PMCID: PMC3752698
- DOI: 10.1038/ni.2670
The scavenger receptor SCARF1 mediates the clearance of apoptotic cells and prevents autoimmunity
Abstract
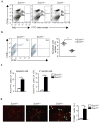
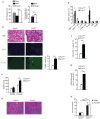
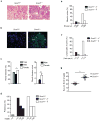
The clearance of apoptotic cells is critical for the control of tissue homeostasis; however, the full range of receptors on phagocytes responsible for the recognition of apoptotic cells remains to be identified. Here we found that dendritic cells (DCs), macrophages and endothelial cells used the scavenger receptor SCARF1 to recognize and engulf apoptotic cells via the complement component C1q. Loss of SCARF1 impaired the uptake of apoptotic cells. Consequently, in SCARF1-deficient mice, dying cells accumulated in tissues, which led to a lupus-like disease, with the spontaneous generation of autoantibodies to DNA-containing antigens, activation of cells of the immune system, dermatitis and nephritis. The discovery of such interactions of SCARF1 with C1q and apoptotic cells provides insight into the molecular mechanisms involved in the maintenance of tolerance and prevention of autoimmune disease.
Conflict of interest statement
COMPETING FINANCIAL INTERESTS
The authors declare no competing financial interests.
Figures
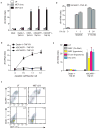
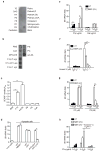
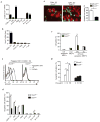
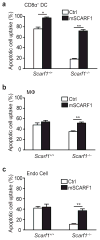

Comment in
-
The danger of sex and death in Scarf1(-/-) autoimmune mice.Nat Immunol. 2013 Sep;14(9):888-9. doi: 10.1038/ni.2684. Nat Immunol. 2013. PMID: 23959178 No abstract available.
Similar articles
-
SCARF1-Induced Efferocytosis Plays an Immunomodulatory Role in Humans, and Autoantibodies Targeting SCARF1 Are Produced in Patients with Systemic Lupus Erythematosus.J Immunol. 2022 Feb 15;208(4):955-967. doi: 10.4049/jimmunol.2100532. Epub 2022 Jan 26. J Immunol. 2022. PMID: 35082161 Free PMC article.
-
The danger of sex and death in Scarf1(-/-) autoimmune mice.Nat Immunol. 2013 Sep;14(9):888-9. doi: 10.1038/ni.2684. Nat Immunol. 2013. PMID: 23959178 No abstract available.
-
Molecular and Cellular Interactions of Scavenger Receptor SR-F1 With Complement C1q Provide Insights Into Its Role in the Clearance of Apoptotic Cells.Front Immunol. 2020 Mar 31;11:544. doi: 10.3389/fimmu.2020.00544. eCollection 2020. Front Immunol. 2020. PMID: 32296440 Free PMC article.
-
SCARF1: a multifaceted, yet largely understudied, scavenger receptor.Inflamm Res. 2018 Aug;67(8):627-632. doi: 10.1007/s00011-018-1154-7. Epub 2018 May 3. Inflamm Res. 2018. PMID: 29725698 Free PMC article. Review.
-
C1q knock-out mice for the study of complement deficiency in autoimmune disease.Exp Clin Immunogenet. 1998;15(4):231-4. doi: 10.1159/000019076. Exp Clin Immunogenet. 1998. PMID: 10072632 Review.
Cited by
-
Scavenger Receptors: Emerging Roles in Cancer Biology and Immunology.Adv Cancer Res. 2015;128:309-64. doi: 10.1016/bs.acr.2015.04.004. Epub 2015 Jun 17. Adv Cancer Res. 2015. PMID: 26216637 Free PMC article. Review.
-
More Than Just a Removal Service: Scavenger Receptors in Leukocyte Trafficking.Front Immunol. 2018 Dec 12;9:2904. doi: 10.3389/fimmu.2018.02904. eCollection 2018. Front Immunol. 2018. PMID: 30631321 Free PMC article. Review.
-
Efferocytosis and autoimmune disease.Int Immunol. 2018 Nov 14;30(12):551-558. doi: 10.1093/intimm/dxy055. Int Immunol. 2018. PMID: 30165442 Free PMC article. Review.
-
Exploring the etiopathogenesis of systemic lupus erythematosus: a genetic perspective.Immunogenetics. 2019 Apr;71(4):283-297. doi: 10.1007/s00251-019-01103-2. Epub 2019 Jan 22. Immunogenetics. 2019. PMID: 30671674 Review.
-
Do follicular dendritic cells regulate lupus-specific B cells?Mol Immunol. 2014 Dec;62(2):283-8. doi: 10.1016/j.molimm.2014.02.010. Epub 2014 Mar 14. Mol Immunol. 2014. PMID: 24636642 Free PMC article. Review.
References
-
- Devitt A, Marshall LJ. The innate immune system and the clearance of apoptotic cells. J Leukoc Biol. 2011;90:447–457. - PubMed
-
- Lauber K, Blumenthal SG, Waibel M, Wesselborg S. Clearance of apoptotic cells: getting rid of the corpses. Mol Cell. 2004;14:277–287. - PubMed
-
- Erwig LP, Henson PM. Clearance of apoptotic cells by phagocytes. Cell death and differentiation. 2008;15:243–250. - PubMed
-
- Munoz LE, et al. Apoptosis in the pathogenesis of systemic lupus erythematosus. Lupus. 2008;17:371–375. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

