CRISPR RNA-guided activation of endogenous human genes
- PMID: 23892898
- PMCID: PMC3794058
- DOI: 10.1038/nmeth.2598
CRISPR RNA-guided activation of endogenous human genes
Abstract
Short guide RNAs (gRNAs) can direct catalytically inactive CRISPR-associated 9 nuclease (dCas9) to repress endogenous genes in bacteria and human cells. Here we show that single or multiple gRNAs can direct dCas9 fused to a VP64 transcriptional activation domain to increase expression of endogenous human genes. This proof-of-principle work shows that clustered regularly interspaced short palindromic repeat (CRISPR)-Cas systems can target heterologous effector domains to endogenous sites in human cells.
Conflict of interest statement
M.L.M. and J.K.J. are inventors on a patent application describing the dCas9-VP64 fusion protein and its use to activate gene expression. J.K.J. has a financial interest in Transposagen Biopharmaceuticals. J.K.J.’s interests were reviewed and are managed by Massachusetts General Hospital and Partners HealthCare in accordance with their conflict of interest policies.
Figures

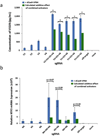
References
-
- Wiedenheft B, Sternberg SH, Doudna JA. RNA-guided genetic silencing systems in bacteria and archaea. Nature. 2012;482:331–338. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

