Clustering and protein dynamics of Drosophila melanogaster telomeres
- PMID: 23893488
- PMCID: PMC3781967
- DOI: 10.1534/genetics.113.155408
Clustering and protein dynamics of Drosophila melanogaster telomeres
Abstract
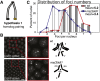
Telomeres are obligatory chromosomal landmarks that demarcate the ends of linear chromosomes to distinguish them from broken ends and can also serve to organize the genome. In both budding and fission yeast, they cluster at the periphery of the nucleus, potentially to establish a compartment of silent chromatin. To gain insight into telomere organization in higher organisms, we investigated their distribution in interphase nuclei of Drosophila melanogaster. We focused on the syncytial blastoderm, an excellent developmental stage for live imaging due to the synchronous division of the nuclei at this time. We followed the EGFP-labeled telomeric protein HOAP in vivo and found that the 16 telomeres yield four to six foci per nucleus, indicative of clustering. Furthermore, we confirmed clustering in other somatic tissues. Importantly, we observed that HOAP signal intensity in the clusters increases in interphase, potentially due to loading of HOAP to newly replicated telomeres. To determine the rules governing clustering, we used in vivo imaging and fluorescence in situ hybridization to test several predictions. First, we inspected mutant embryos that develop as haploids and found that clustering is not mediated by associations between homologs. Second, we probed specifically for a telomere of novel sequence and found strong evidence against DNA sequence identity and homology as critical factors. Third, we ruled out predominance of intrachromosomal interactions by marking both ends of a chromosome. Based on these results, we propose that clustering is independent of sequence and is likely maintained by an as yet undetermined factor.
Keywords: Drosophila telomeres; nuclear organization; telomere; telomere clustering; telomere dynamics.
Figures






Similar articles
-
HipHop interacts with HOAP and HP1 to protect Drosophila telomeres in a sequence-independent manner.EMBO J. 2010 Feb 17;29(4):819-29. doi: 10.1038/emboj.2009.394. Epub 2010 Jan 7. EMBO J. 2010. PMID: 20057353 Free PMC article.
-
The Drosophila modigliani (moi) gene encodes a HOAP-interacting protein required for telomere protection.Proc Natl Acad Sci U S A. 2009 Feb 17;106(7):2271-6. doi: 10.1073/pnas.0812702106. Epub 2009 Jan 30. Proc Natl Acad Sci U S A. 2009. PMID: 19181850 Free PMC article.
-
Epigenetic maintenance of telomere identity in Drosophila: buckle up for the sperm ride.Cell Cycle. 2011 Apr 1;10(7):1037-42. doi: 10.4161/cc.10.7.15071. Epub 2011 Apr 1. Cell Cycle. 2011. PMID: 21386659
-
The mechanism of telomere protection: a comparison between Drosophila and humans.Chromosoma. 2005 Aug;114(3):135-45. doi: 10.1007/s00412-005-0005-9. Epub 2005 Jul 13. Chromosoma. 2005. PMID: 16012858 Review.
-
Terminin: a protein complex that mediates epigenetic maintenance of Drosophila telomeres.Nucleus. 2011 Sep-Oct;2(5):383-91. doi: 10.4161/nucl.2.5.17873. Epub 2011 Sep 1. Nucleus. 2011. PMID: 21989238 Review.
Cited by
-
Protection of Drosophila chromosome ends through minimal telomere capping.J Cell Sci. 2015 May 15;128(10):1969-81. doi: 10.1242/jcs.167825. Epub 2015 Apr 23. J Cell Sci. 2015. PMID: 25908850 Free PMC article.
-
Key role of piRNAs in telomeric chromatin maintenance and telomere nuclear positioning in Drosophila germline.Epigenetics Chromatin. 2018 Jul 12;11(1):40. doi: 10.1186/s13072-018-0210-4. Epigenetics Chromatin. 2018. PMID: 30001204 Free PMC article.
-
The nanoCUT&RUN technique visualizes telomeric chromatin in Drosophila.PLoS Genet. 2022 Sep 1;18(9):e1010351. doi: 10.1371/journal.pgen.1010351. eCollection 2022 Sep. PLoS Genet. 2022. PMID: 36048878 Free PMC article.
-
MTV, an ssDNA Protecting Complex Essential for Transposon-Based Telomere Maintenance in Drosophila.PLoS Genet. 2016 Nov 11;12(11):e1006435. doi: 10.1371/journal.pgen.1006435. eCollection 2016 Nov. PLoS Genet. 2016. PMID: 27835648 Free PMC article.
-
Specific Localization of the Drosophila Telomere Transposon Proteins and RNAs, Give Insight in Their Behavior, Control and Telomere Biology in This Organism.PLoS One. 2015 Jun 12;10(6):e0128573. doi: 10.1371/journal.pone.0128573. eCollection 2015. PLoS One. 2015. PMID: 26068215 Free PMC article.
References
-
- Bi X., Wei S. C., Rong Y. S., 2004. Telomere protection without a telomerase; the role of ATM and Mre11 in Drosophila telomere maintenance. Curr. Biol. 14: 1348–1353. - PubMed
-
- Billia F., De Boni U., 1991. Localization of centromeric satellite and telomeric DNA sequences in dorsal root ganglion neurons, in vitro. J. Cell Sci. 100(Pt 1): 219–226. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

