Genome-wide upstream motif analysis of Cryptosporidium parvum genes clustered by expression profile
- PMID: 23895416
- PMCID: PMC3734150
- DOI: 10.1186/1471-2164-14-516
Genome-wide upstream motif analysis of Cryptosporidium parvum genes clustered by expression profile
Abstract
Background: There are very few molecular genetic tools available to study the apicomplexan parasite Cryptosporidium parvum. The organism is not amenable to continuous in vitro cultivation or transfection, and purification of intracellular developmental stages in sufficient numbers for most downstream molecular applications is difficult and expensive since animal hosts are required. As such, very little is known about gene regulation in C. parvum.
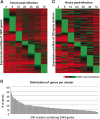
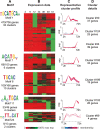
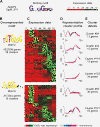
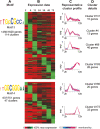
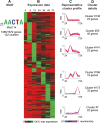
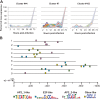
Results: We have clustered whole-genome gene expression profiles generated from a previous study of seven post-infection time points of 3,281 genes to identify genes that show similar expression patterns throughout the first 72 hours of in vitro epithelial cell culture. We used the algorithms MEME, AlignACE and FIRE to identify conserved, overrepresented DNA motifs in the upstream promoter region of genes with similar expression profiles. The most overrepresented motifs were E2F (5'-TGGCGCCA-3'); G-box (5'-G.GGGG-3'); a well-documented ApiAP2 binding motif (5'-TGCAT-3'), and an unknown motif (5'-[A/C] AACTA-3'). We generated a recombinant C. parvum DNA-binding protein domain from a putative ApiAP2 transcription factor [CryptoDB: cgd8_810] and determined its binding specificity using protein-binding microarrays. We demonstrate that cgd8_810 can putatively bind the overrepresented G-box motif, implicating this ApiAP2 in the regulation of many gene clusters.
Conclusion: Several DNA motifs were identified in the upstream sequences of gene clusters that might serve as potential cis-regulatory elements. These motifs, in concert with protein DNA binding site data, establish for the first time the beginnings of a global C. parvum gene regulatory map that will contribute to our understanding of the development of this zoonotic parasite.
Figures


Similar articles
-
The Cryptosporidium parvum ApiAP2 gene family: insights into the evolution of apicomplexan AP2 regulatory systems.Nucleic Acids Res. 2014 Jul;42(13):8271-84. doi: 10.1093/nar/gku500. Epub 2014 Jun 23. Nucleic Acids Res. 2014. PMID: 24957599 Free PMC article.
-
Identification of putative cis-regulatory elements in Cryptosporidium parvum by de novo pattern finding.BMC Genomics. 2007 Jan 9;8:13. doi: 10.1186/1471-2164-8-13. BMC Genomics. 2007. PMID: 17212834 Free PMC article.
-
A highly divergent 33 kDa Cryptosporidium parvum antigen.J Parasitol. 2014 Aug;100(4):527-31. doi: 10.1645/13-433.1. Epub 2014 Mar 6. J Parasitol. 2014. PMID: 24601821
-
Cryptosporidium parvum gene discovery.Adv Exp Med Biol. 1999;473:241-7. doi: 10.1007/978-1-4615-4143-1_26. Adv Exp Med Biol. 1999. PMID: 10659365 Review.
-
Reduction and compaction in the genome of the apicomplexan parasite Cryptosporidium parvum.Dev Cell. 2004 May;6(5):614-6. doi: 10.1016/s1534-5807(04)00135-2. Dev Cell. 2004. PMID: 15130487 Review.
Cited by
-
ApiAP2 Factors as Candidate Regulators of Stochastic Commitment to Merozoite Production in Theileria annulata.PLoS Negl Trop Dis. 2015 Aug 14;9(8):e0003933. doi: 10.1371/journal.pntd.0003933. eCollection 2015. PLoS Negl Trop Dis. 2015. PMID: 26273826 Free PMC article.
-
Gene regulation in Cryptosporidium: New insights and unanswered questions.Curr Res Parasitol Vector Borne Dis. 2025 Jun 17;8:100280. doi: 10.1016/j.crpvbd.2025.100280. eCollection 2025. Curr Res Parasitol Vector Borne Dis. 2025. PMID: 40656160 Free PMC article.
-
Genetic Ablation of a Female-Specific Apetala 2 Transcription Factor Blocks Oocyst Shedding in Cryptosporidium parvum.mBio. 2023 Apr 25;14(2):e0326122. doi: 10.1128/mbio.03261-22. Epub 2023 Feb 14. mBio. 2023. PMID: 36786597 Free PMC article.
-
Past and future trends of Cryptosporidium in vitro research.Exp Parasitol. 2019 Jan;196:28-37. doi: 10.1016/j.exppara.2018.12.001. Epub 2018 Dec 3. Exp Parasitol. 2019. PMID: 30521793 Free PMC article. Review.
-
The Cryptosporidium parvum ApiAP2 gene family: insights into the evolution of apicomplexan AP2 regulatory systems.Nucleic Acids Res. 2014 Jul;42(13):8271-84. doi: 10.1093/nar/gku500. Epub 2014 Jun 23. Nucleic Acids Res. 2014. PMID: 24957599 Free PMC article.
References
-
- Kotloff KL, Nataro JP, Blackwelder WC, Nasrin D, Farag TH, Panchalingam S, Wu Y, Sow SO, Sur D, Breiman RF, Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case–control study. Lancet. 2013. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

