Erythropoietic defect associated with reduced cell proliferation in mice lacking the 26S proteasome shuttling factor Rad23b
- PMID: 23897431
- PMCID: PMC3811862
- DOI: 10.1128/MCB.05772-11
Erythropoietic defect associated with reduced cell proliferation in mice lacking the 26S proteasome shuttling factor Rad23b
Abstract
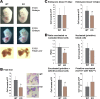
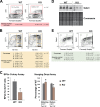
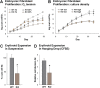
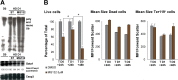
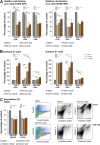
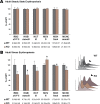
Rad23a and Rad23b proteins are linked to nucleotide excision DNA repair (NER) via association with the DNA damage recognition protein xeroderma pigmentosum group C (XPC) are and known to be implicated in protein turnover by the 26S proteasome. Rad23b-null mice are NER proficient, likely due to the redundant function of the Rad23b paralogue, Rad23a. However, Rad23b-null midgestation embryos are anemic, and most embryos die before birth. Using an unbiased proteomics approach, we found that the majority of Rad23b-interacting partners are associated with the ubiquitin-proteasome system (UPS). We tested the requirement for Rad23b-dependent UPS activity in cellular proliferation and more specifically in the process of erythropoiesis. In cultured fibroblasts derived from embryos lacking Rad23b, proliferation rates were reduced. In fetal livers of Rad23b-null embryos, we observed reduced proliferation, accumulation of early erythroid progenitors, and a block during erythroid maturation. In primary wild-type (WT) erythroid cells, knockdown of Rad23b or chemical inhibition of the proteasome reduced survival and differentiation capability. Finally, the defects linked to Rad23b loss specifically affected fetal definitive erythropoiesis and stress erythropoiesis in adult mice. Together, these data indicate a previously unappreciated requirement for Rad23b and the UPS in regulation of proliferation in different cell types.
Figures


Similar articles
-
The RAD23 family provides an essential connection between the 26S proteasome and ubiquitylated proteins in Arabidopsis.Plant Cell. 2010 Jan;22(1):124-42. doi: 10.1105/tpc.109.072660. Epub 2010 Jan 19. Plant Cell. 2010. PMID: 20086187 Free PMC article.
-
Human granulocyte-macrophage colony-stimulating factor (hGM-CSF) stimulates primitive and definitive erythropoiesis in mouse embryos expressing hGM-CSF receptors but not erythropoietin receptors.Blood. 2001 Dec 15;98(13):3618-25. doi: 10.1182/blood.v98.13.3618. Blood. 2001. PMID: 11739165
-
Arabidopsis RAD23B regulates pollen development by mediating degradation of KRP1.J Exp Bot. 2020 Jul 6;71(14):4010-4019. doi: 10.1093/jxb/eraa167. J Exp Bot. 2020. PMID: 32242227
-
Molecular basis for damage recognition and verification by XPC-RAD23B and TFIIH in nucleotide excision repair.DNA Repair (Amst). 2018 Nov;71:33-42. doi: 10.1016/j.dnarep.2018.08.005. Epub 2018 Aug 23. DNA Repair (Amst). 2018. PMID: 30174301 Free PMC article. Review.
-
Primitive erythropoiesis in the mammalian embryo.Int J Dev Biol. 2010;54(6-7):1011-8. doi: 10.1387/ijdb.093056jp. Int J Dev Biol. 2010. PMID: 20711979 Review.
Cited by
-
Functional and mechanistic studies of XPC DNA-repair complex as transcriptional coactivator in embryonic stem cells.Proc Natl Acad Sci U S A. 2015 May 5;112(18):E2317-26. doi: 10.1073/pnas.1505569112. Epub 2015 Apr 21. Proc Natl Acad Sci U S A. 2015. PMID: 25901318 Free PMC article.
-
Damage-Net: A program for DNA repair meta-analysis identifies a network of novel repair genes that facilitate cancer evolution.DNA Repair (Amst). 2021 Sep;105:103158. doi: 10.1016/j.dnarep.2021.103158. Epub 2021 Jun 10. DNA Repair (Amst). 2021. PMID: 34147942 Free PMC article.
References
-
- Dantuma NP, Heinen C, Hoogstraten D. 2009. The ubiquitin receptor Rad23: at the crossroads of nucleotide excision repair and proteasomal degradation. DNA Repair (Amst.) 8:449–460 - PubMed
-
- Hiyama H, Yokoi M, Masutani C, Sugasawa K, Maekawa T, Tanaka K, Hoeijmakers JH, Hanaoka F. 1999. Interaction of hHR23 with S5a. The ubiquitin-like domain of hHR23 mediates interaction with S5a subunit of 26 S proteasome. J. Biol. Chem. 274:28019–28025 - PubMed
-
- van der Spek PJ, Visser CE, Hanaoka F, Smit B, Hagemeijer A, Bootsma D, Hoeijmakers JH. 1996. Cloning, comparative mapping, and RNA expression of the mouse homologues of the Saccharomyces cerevisiae nucleotide excision repair gene RAD23. Genomics 31:20–27 - PubMed
-
- Miller RD, Prakash L, Prakash S. 1982. Defective excision of pyrimidine dimers and interstrand DNA crosslinks in rad7 and rad23 mutants of Saccharomyces cerevisiae. Mol. Gen. Genet. 188:235–239 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
