DNA structure-specific priming of ATR activation by DNA-PKcs
- PMID: 23897887
- PMCID: PMC3734074
- DOI: 10.1083/jcb.201304139
DNA structure-specific priming of ATR activation by DNA-PKcs
Abstract
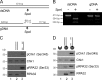
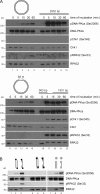
Three phosphatidylinositol-3-kinase-related protein kinases implement cellular responses to DNA damage. DNA-dependent protein kinase catalytic subunit (DNA-PKcs) and ataxia-telangiectasia mutated respond primarily to DNA double-strand breaks (DSBs). Ataxia-telangiectasia and RAD3-related (ATR) signals the accumulation of replication protein A (RPA)-covered single-stranded DNA (ssDNA), which is caused by replication obstacles. Stalled replication intermediates can further degenerate and yield replication-associated DSBs. In this paper, we show that the juxtaposition of a double-stranded DNA end and a short ssDNA gap triggered robust activation of endogenous ATR and Chk1 in human cell-free extracts. This DNA damage signal depended on DNA-PKcs and ATR, which congregated onto gapped linear duplex DNA. DNA-PKcs primed ATR/Chk1 activation through DNA structure-specific phosphorylation of RPA32 and TopBP1. The synergistic activation of DNA-PKcs and ATR suggests that the two kinases combine to mount a prompt and specific response to replication-born DSBs.
Figures



Similar articles
-
DNA-PKcs is required to maintain stability of Chk1 and Claspin for optimal replication stress response.Nucleic Acids Res. 2014 Apr;42(7):4463-73. doi: 10.1093/nar/gku116. Epub 2014 Feb 5. Nucleic Acids Res. 2014. PMID: 24500207 Free PMC article.
-
Distinct roles for DNA-PK, ATM and ATR in RPA phosphorylation and checkpoint activation in response to replication stress.Nucleic Acids Res. 2012 Nov;40(21):10780-94. doi: 10.1093/nar/gks849. Epub 2012 Sep 12. Nucleic Acids Res. 2012. PMID: 22977173 Free PMC article.
-
ATR-dependent phosphorylation of DNA-dependent protein kinase catalytic subunit in response to UV-induced replication stress.Mol Cell Biol. 2006 Oct;26(20):7520-8. doi: 10.1128/MCB.00048-06. Epub 2006 Aug 14. Mol Cell Biol. 2006. PMID: 16908529 Free PMC article.
-
DNA-Dependent Protein Kinase Catalytic Subunit: The Sensor for DNA Double-Strand Breaks Structurally and Functionally Related to Ataxia Telangiectasia Mutated.Genes (Basel). 2021 Jul 27;12(8):1143. doi: 10.3390/genes12081143. Genes (Basel). 2021. PMID: 34440313 Free PMC article. Review.
-
Activation of ATR-related protein kinase upon DNA damage recognition.Curr Genet. 2020 Apr;66(2):327-333. doi: 10.1007/s00294-019-01039-w. Epub 2019 Oct 17. Curr Genet. 2020. PMID: 31624858 Free PMC article. Review.
Cited by
-
The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA.Nat Commun. 2024 Feb 29;15(1):1892. doi: 10.1038/s41467-024-46208-9. Nat Commun. 2024. PMID: 38424107 Free PMC article.
-
RPA-coated single-stranded DNA as a platform for post-translational modifications in the DNA damage response.Cell Res. 2015 Jan;25(1):9-23. doi: 10.1038/cr.2014.147. Epub 2014 Nov 18. Cell Res. 2015. PMID: 25403473 Free PMC article. Review.
-
DNA-PKcs is required to maintain stability of Chk1 and Claspin for optimal replication stress response.Nucleic Acids Res. 2014 Apr;42(7):4463-73. doi: 10.1093/nar/gku116. Epub 2014 Feb 5. Nucleic Acids Res. 2014. PMID: 24500207 Free PMC article.
-
mTORC1 and DNA-PKcs as novel molecular determinants of sensitivity to Chk1 inhibition.Mol Oncol. 2016 Jan;10(1):101-12. doi: 10.1016/j.molonc.2015.08.004. Epub 2015 Aug 25. Mol Oncol. 2016. PMID: 26471831 Free PMC article.
-
Working on Genomic Stability: From the S-Phase to Mitosis.Genes (Basel). 2020 Feb 20;11(2):225. doi: 10.3390/genes11020225. Genes (Basel). 2020. PMID: 32093406 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

