Infection structure-specific expression of β-1,3-glucan synthase is essential for pathogenicity of Colletotrichum graminicola and evasion of β-glucan-triggered immunity in maize
- PMID: 23898035
- PMCID: PMC3723631
- DOI: 10.1105/tpc.112.103499
Infection structure-specific expression of β-1,3-glucan synthase is essential for pathogenicity of Colletotrichum graminicola and evasion of β-glucan-triggered immunity in maize
Abstract
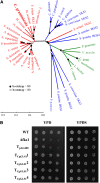
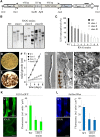
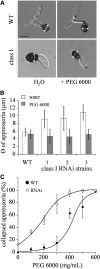
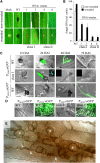
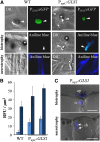
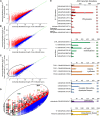
β-1,3-Glucan and chitin are the most prominent polysaccharides of the fungal cell wall. Covalently linked, these polymers form a scaffold that determines the form and properties of vegetative and pathogenic hyphae. While the role of chitin in plant infection is well understood, the role of β-1,3-glucan is unknown. We functionally characterized the β-1,3-glucan synthase gene GLS1 of the maize (Zea mays) pathogen Colletotrichum graminicola, employing RNA interference (RNAi), GLS1 overexpression, live-cell imaging, and aniline blue fluorochrome staining. This hemibiotroph sequentially differentiates a melanized appressorium on the cuticle and biotrophic and necrotrophic hyphae in its host. Massive β-1,3-glucan contents were detected in cell walls of appressoria and necrotrophic hyphae. Unexpectedly, GLS1 expression and β-1,3-glucan contents were drastically reduced during biotrophic development. In appressoria of RNAi strains, downregulation of β-1,3-glucan synthesis increased cell wall elasticity, and the appressoria exploded. While the shape of biotrophic hyphae was unaffected in RNAi strains, necrotrophic hyphae showed severe distortions. Constitutive expression of GLS1 led to exposure of β-1,3-glucan on biotrophic hyphae, massive induction of broad-spectrum defense responses, and significantly reduced disease symptom severity. Thus, while β-1,3-glucan synthesis is required for cell wall rigidity in appressoria and fast-growing necrotrophic hyphae, its rigorous downregulation during biotrophic development represents a strategy for evading β-glucan-triggered immunity.
Figures
References
-
- Afzal A.J., Wood A.J., Lightfoot D.A. (2008). Plant receptor-like serine threonine kinases: Roles in signaling and plant defense. Mol. Plant Microbe Interact. 21: 507–517 - PubMed
-
- Aimanianda V., Bayry J., Bozza S., Kniemeyer O., Perruccio K., Elluru S.R., Clavaud C., Paris S., Brakhage A.A., Kaveri S.V., Romani L., Latgé J.-P. (2009). Surface hydrophobin prevents immune recognition of airborne fungal spores. Nature 460: 1117–1121 - PubMed
-
- Almagro L., Gómez Ros L.V., Belchi-Navarro S., Bru R., Ros Barceló A., Pedreño M.A. (2009). Class III peroxidases in plant defence reactions. J. Exp. Bot. 60: 377–390 - PubMed
-
- Andrie R.M., Martinez J.P., Ciuffetti L.M. (2005). Development of ToxA and ToxB promoter-driven fluorescent protein expression vectors for use in filamentous ascomycetes. Mycologia 97: 1152–1161 - PubMed
-
- Barber M.S., Bertram R.E., Ride J.P. (1989). Chitin oligosaccharides elicit lignification in wounded wheat leaves. Physiol. Mol. Plant Pathol. 34: 3–12
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

