Structural basis for the stabilization of the complement alternative pathway C3 convertase by properdin
- PMID: 23901101
- PMCID: PMC3746899
- DOI: 10.1073/pnas.1309618110
Structural basis for the stabilization of the complement alternative pathway C3 convertase by properdin
Abstract
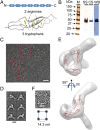
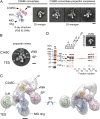
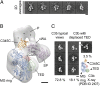
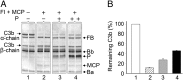
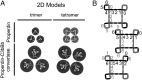
Complement is an essential component of innate immunity. Its activation results in the assembly of unstable protease complexes, denominated C3/C5 convertases, leading to inflammation and lysis. Regulatory proteins inactivate C3/C5 convertases on host surfaces to avoid collateral tissue damage. On pathogen surfaces, properdin stabilizes C3/C5 convertases to efficiently fight infection. How properdin performs this function is, however, unclear. Using electron microscopy we show that the N- and C-terminal ends of adjacent monomers in properdin oligomers conform a curly vertex that holds together the AP convertase, interacting with both the C345C and vWA domains of C3b and Bb, respectively. Properdin also promotes a large displacement of the TED (thioester-containing domain) and CUB (complement protein subcomponents C1r/C1s, urchin embryonic growth factor and bone morphogenetic protein 1) domains of C3b, which likely impairs C3-convertase inactivation by regulatory proteins. The combined effect of molecular cross-linking and structural reorganization increases stability of the C3 convertase and facilitates recruitment of fluid-phase C3 convertase to the cell surfaces. Our model explains how properdin mediates the assembly of stabilized C3/C5-convertase clusters, which helps to localize complement amplification to pathogen surfaces.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Rodríguez de Córdoba S, Harris CL, Morgan BP, Llorca O. Lessons from functional and structural analyses of disease-associated genetic variants in the complement alternative pathway. Biochim Biophys Acta. 2011;1812(1):12–22. - PubMed
-
- Gros P, Milder FJ, Janssen BJ. Complement driven by conformational changes. Nat Rev Immunol. 2008;8(1):48–58. - PubMed
-
- Hourcade DE. The role of properdin in the assembly of the alternative pathway C3 convertases of complement. J Biol Chem. 2006;281(4):2128–2132. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

