WHSC1 links transcription elongation to HIRA-mediated histone H3.3 deposition
- PMID: 23921552
- PMCID: PMC3770338
- DOI: 10.1038/emboj.2013.176
WHSC1 links transcription elongation to HIRA-mediated histone H3.3 deposition
Abstract
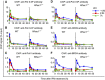
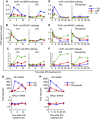
Actively transcribed genes are enriched with the histone variant H3.3. Although H3.3 deposition has been linked to transcription, mechanisms controlling this process remain elusive. We investigated the role of the histone methyltransferase Wolf-Hirschhorn syndrome candidate 1 (WHSC1) (NSD2/MMSET) in H3.3 deposition into interferon (IFN) response genes. IFN treatment triggered robust H3.3 incorporation into activated genes, which continued even after cessation of transcription. Likewise, UV radiation caused H3.3 deposition in UV-activated genes. However, in Whsc1(-/-) cells IFN- or UV-triggered H3.3 deposition was absent, along with a marked reduction in IFN- or UV-induced transcription. We found that WHSC1 interacted with the bromodomain protein 4 (BRD4) and the positive transcription elongation factor b (P-TEFb) and facilitated transcriptional elongation. WHSC1 also associated with HIRA, the H3.3-specific histone chaperone, independent of BRD4 and P-TEFb. WHSC1 and HIRA co-occupied IFN-stimulated genes and supported prolonged H3.3 incorporation, leaving a lasting transcriptional mark. Our results reveal a previously unrecognized role of WHSC1, which links transcriptional elongation and H3.3 deposition into activated genes through two molecularly distinct pathways.
Conflict of interest statement
The authors declared that they have no conflict of interests.
Figures







Similar articles
-
HIRA vs. DAXX: the two axes shaping the histone H3.3 landscape.Exp Mol Med. 2024 Feb;56(2):251-263. doi: 10.1038/s12276-023-01145-3. Epub 2024 Feb 1. Exp Mol Med. 2024. PMID: 38297159 Free PMC article. Review.
-
H3.Y discriminates between HIRA and DAXX chaperone complexes and reveals unexpected insights into human DAXX-H3.3-H4 binding and deposition requirements.Nucleic Acids Res. 2017 Jun 2;45(10):5691-5706. doi: 10.1093/nar/gkx131. Nucleic Acids Res. 2017. PMID: 28334823 Free PMC article.
-
A histone H3 lysine 36 trimethyltransferase links Nkx2-5 to Wolf-Hirschhorn syndrome.Nature. 2009 Jul 9;460(7252):287-91. doi: 10.1038/nature08086. Epub 2009 May 31. Nature. 2009. PMID: 19483677
-
Intellectual disability-associated dBRWD3 regulates gene expression through inhibition of HIRA/YEM-mediated chromatin deposition of histone H3.3.EMBO Rep. 2015 Apr;16(4):528-38. doi: 10.15252/embr.201439092. Epub 2015 Feb 9. EMBO Rep. 2015. PMID: 25666827 Free PMC article.
-
A Molecular Prospective for HIRA Complex Assembly and H3.3-Specific Histone Chaperone Function.J Mol Biol. 2017 Jun 30;429(13):1924-1933. doi: 10.1016/j.jmb.2016.11.010. Epub 2016 Nov 19. J Mol Biol. 2017. PMID: 27871933 Free PMC article. Review.
Cited by
-
Establishment and function of chromatin modification at enhancers.Open Biol. 2020 Oct;10(10):200255. doi: 10.1098/rsob.200255. Epub 2020 Oct 14. Open Biol. 2020. PMID: 33050790 Free PMC article.
-
HIRA vs. DAXX: the two axes shaping the histone H3.3 landscape.Exp Mol Med. 2024 Feb;56(2):251-263. doi: 10.1038/s12276-023-01145-3. Epub 2024 Feb 1. Exp Mol Med. 2024. PMID: 38297159 Free PMC article. Review.
-
Super Enhancer-Mediated Upregulation of HJURP Promotes Growth and Survival of t(4;14)-Positive Multiple Myeloma.Cancer Res. 2022 Feb 1;82(3):406-418. doi: 10.1158/0008-5472.CAN-21-0921. Epub 2021 Dec 10. Cancer Res. 2022. PMID: 34893510 Free PMC article.
-
Wolf-Hirschhorn Syndrome Candidate 1 Is Necessary for Correct Hematopoietic and B Cell Development.Cell Rep. 2017 May 23;19(8):1586-1601. doi: 10.1016/j.celrep.2017.04.069. Cell Rep. 2017. PMID: 28538178 Free PMC article.
-
Live cell analysis of mobility and decay kinetics of the histone variant H3.3.J Biol Chem. 2025 Jun;301(6):108557. doi: 10.1016/j.jbc.2025.108557. Epub 2025 Apr 30. J Biol Chem. 2025. PMID: 40316020 Free PMC article.
References
-
- Bergemann AD, Cole F, Hirschhorn K (2005) The etiology of Wolf-Hirschhorn syndrome. Trends Genet 21: 188–195 - PubMed
-
- Carrozza MJ, Li B, Florens L, Suganuma T, Swanson SK, Lee KK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005) Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123: 581–592 - PubMed
-
- Chao SH, Fujinaga K, Marion JE, Taube R, Sausville EA, Senderowicz AM, Peterlin BM, Price DH (2000) Flavopiridol inhibits P-TEFb and blocks HIV-1 replication. J Biol Chem 275: 28345–28348 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

