Aggregation of lipid-anchored full-length H-Ras in lipid bilayers: simulations with the MARTINI force field
- PMID: 23923044
- PMCID: PMC3724741
- DOI: 10.1371/journal.pone.0071018
Aggregation of lipid-anchored full-length H-Ras in lipid bilayers: simulations with the MARTINI force field
Abstract
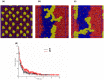
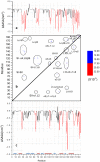
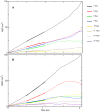
Lipid-anchored Ras oncoproteins assemble into transient, nano-sized substructures on the plasma membrane. These substructures, called nanoclusters, were proposed to be crucial for high-fidelity signal transmission in cells. However, the molecular basis of Ras nanoclustering is poorly understood. In this work, we used coarse-grained (CG) molecular dynamics simulations to investigate the molecular mechanism by which full-length H-ras proteins form nanoclusters in a model membrane. We chose two different conformations of H-ras that were proposed to represent the active and inactive state of the protein, and a domain-forming model bilayer made up of di16:0-PC (DPPC), di18:2-PC (DLiPC) and cholesterol. We found that, irrespective of the initial conformation, Ras molecules assembled into a single large aggregate. However, the two binding modes, which are characterized by the different orientation of the G-domain with respect to the membrane, differ in dynamics and organization during and after aggregation. Some of these differences involve regions of Ras that are important for effector/modulator binding, which may partly explain observed differences in the ability of active and inactive H-ras nanoclusters to recruit effectors. The simulations also revealed some limitations in the CG force field to study protein assembly in solution, which we discuss in the context of proposed potential avenues of improvement.
Conflict of interest statement
Figures



References
-
- Barbacid M (1987) ras Genes. Annual Review of Biochemistry 56: 779–827. - PubMed
-
- Wennerberg K, Rossman KL, Der CJ (2005) The Ras superfamily at a glance. Journal of Cell Science 118: 843–846. - PubMed
-
- Yan J, Roy S, Apolloni A, Lane A, Hancock JF (1998) Ras Isoforms Vary in Their Ability to Activate Raf-1 and Phosphoinositide 3-Kinase. Journal of Biological Chemistry 273: 24052–24056. - PubMed
-
- Voice JK, Klemke RL, Le A, Jackson JH (1999) Four Human Ras Homologs Differ in Their Abilities to Activate Raf-1, Induce Transformation, and Stimulate Cell Motility. Journal of Biological Chemistry 274: 17164–17170. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

