Functional endogenous viral elements in the genome of the parasitoid wasp Cotesia congregata: insights into the evolutionary dynamics of bracoviruses
- PMID: 23938757
- PMCID: PMC3758192
- DOI: 10.1098/rstb.2013.0047
Functional endogenous viral elements in the genome of the parasitoid wasp Cotesia congregata: insights into the evolutionary dynamics of bracoviruses
Abstract
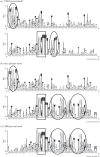
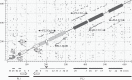
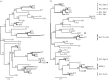
Bracoviruses represent the most complex endogenous viral elements (EVEs) described to date. Nudiviral genes have been hosted within parasitoid wasp genomes since approximately 100 Ma. They play a crucial role in the wasp life cycle as they produce bracovirus particles, which are injected into parasitized lepidopteran hosts during wasp oviposition. Bracovirus particles encapsidate multiple dsDNA circles encoding virulence genes. Their expression in parasitized caterpillars is essential for wasp parasitism success. Here, we report on the genomic organization of the proviral segments (i.e. master sequences used to produce the encapsidated dsDNA circles) present in the Cotesia congregata parasitoid wasp genome. The provirus is composed of a macrolocus, comprising two-thirds of the proviral segments and of seven dispersed loci, each containing one to three segments. Comparative genomic analyses with closely related species gave insights into the evolutionary dynamics of bracovirus genomes. Conserved synteny in the different wasp genomes showed the orthology of the proviral macrolocus across different species. The nudiviral gene odv-e66-like1 is conserved within the macrolocus, suggesting an ancient co-localization of the nudiviral genome and bracovirus proviral segments. By contrast, the evolution of proviral segments within the macrolocus has involved a series of lineage-specific duplications.
Keywords: bracovirus; comparative genomics; obligatory mutualism; parasitoid wasp; polydnavirus.
Figures





References
-
- Herniou EA, Huguet E, Thézé J, Bézier A, Periquet G, Drezen J-M. 2013. When parasitic wasps hijacked viruses: genomic and functional evolution of polydnaviruses. Phil. Trans. R. Soc. B 368, 20130051 (doi:10.1098/rstb.2013.0051) - DOI - PMC - PubMed
-
- Dupuy C, Huguet E, Drezen J-M. 2006. Unfolding the evolutionary story of polydnaviruses. Virus Res. 117, 81–89 (doi:10.1016/j.virusres.2006.01.001) - DOI - PubMed
-
- Murphy N, Banks JC, Whitfield JB, Austin AD. 2008. Phylogeny of the parasitic microgastroid subfamilies (Hymenoptera: Braconidae) based on sequence data from seven genes, with an improved time estimate of the origin of the lineage. Mol. Phylogenet. Evol. 47, 378–395 (doi:10.1016/j.ympev.2008.01.022) - DOI - PubMed
-
- Bézier A, et al. 2009. Polydnaviruses of braconid wasps derive from an ancestral nudivirus. Science 323, 926–930 (doi:10.1126/science.1166788) - DOI - PubMed
-
- Théze J, Bézier A, Periquet G, Drezen J-M, Herniou EA. 2011. Paleozoic origin of insect large dsDNA viruses. Proc. Natl Acad. Sci. USA 108, 15 931–15 935 (doi:10.1073/pnas.1105580108) - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
