Linking stochastic fluctuations in chromatin structure and gene expression
- PMID: 23940458
- PMCID: PMC3735467
- DOI: 10.1371/journal.pbio.1001621
Linking stochastic fluctuations in chromatin structure and gene expression
Abstract
The number of mRNA and protein molecules expressed from a single gene molecule fluctuates over time. These fluctuations have been attributed, in part, to the random transitioning of promoters between transcriptionally active and inactive states, causing transcription to occur in bursts. However, the molecular basis of transcriptional bursting remains poorly understood. By electron microscopy of single PHO5 gene molecules from yeast, we show that the "activated" promoter assumes alternative nucleosome configurations at steady state, including the maximally repressive, fully nucleosomal, and the maximally non-repressive, nucleosome-free, configuration. We demonstrate that the observed probabilities of promoter nucleosome configurations are obtained from a simple, intrinsically stochastic process of nucleosome assembly, disassembly, and position-specific sliding; and we show that gene expression and promoter nucleosome configuration can be mechanistically coupled, relating promoter nucleosome dynamics and gene expression fluctuations. Together, our findings suggest a structural basis for transcriptional bursting, and offer new insights into the mechanism of transcriptional regulation and the kinetics of promoter nucleosome transitions.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
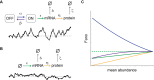
 indicate degradation of the gene product. Greek letters refer to transition probabilities per unit time and molecule (“kinetic parameters”); below, a typical time trace (black curve) for the fluctuation in single cell mRNA molecule number about the steady state mean (dashed gray line). (B) “Deterministic model” of a transcriptionally active gene. The black curve beneath the model represents a typical time trace of mRNA fluctuation about the same mean (dashed gray line) as in (A). (C) Steady-state Fano factor values (Fano) were calculated as a function of a single kinetic parameter (the “regulatory parameter”), with all other kinetic parameters held constant. The coloring of the resulting noise profiles refers to the identity of the kinetic parameter that was allowed to float to vary the mean abundance of protein molecules (mean abundance). Thus, blue refers to the bursting frequency α, see (A), etc. The dashed green line indicates the expected Fano profile for the modulation of ε for the deterministic model B. Noise profiles were determined by analytical calculations as described in Materials and Methods.
indicate degradation of the gene product. Greek letters refer to transition probabilities per unit time and molecule (“kinetic parameters”); below, a typical time trace (black curve) for the fluctuation in single cell mRNA molecule number about the steady state mean (dashed gray line). (B) “Deterministic model” of a transcriptionally active gene. The black curve beneath the model represents a typical time trace of mRNA fluctuation about the same mean (dashed gray line) as in (A). (C) Steady-state Fano factor values (Fano) were calculated as a function of a single kinetic parameter (the “regulatory parameter”), with all other kinetic parameters held constant. The coloring of the resulting noise profiles refers to the identity of the kinetic parameter that was allowed to float to vary the mean abundance of protein molecules (mean abundance). Thus, blue refers to the bursting frequency α, see (A), etc. The dashed green line indicates the expected Fano profile for the modulation of ε for the deterministic model B. Noise profiles were determined by analytical calculations as described in Materials and Methods.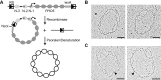


 ; i.e., R was
; i.e., R was  -fold more probable given (D) than given (B) (Materials and Methods). (D) Transition topology with unidirectional nucleosome sliding; dashed arrows indicate sliding transitions. (E) Same as (A), with predictions based on the topology in (F);
-fold more probable given (D) than given (B) (Materials and Methods). (D) Transition topology with unidirectional nucleosome sliding; dashed arrows indicate sliding transitions. (E) Same as (A), with predictions based on the topology in (F);  (and hence
(and hence  ). (G) Transition topology for “stable nucleosome retention.” This hypothesis was disproved by R, for
). (G) Transition topology for “stable nucleosome retention.” This hypothesis was disproved by R, for  , but
, but  ; thus, its likelihood, given R, was
; thus, its likelihood, given R, was  . (H) Transition topology for all-or-nothing disassembly:
. (H) Transition topology for all-or-nothing disassembly:  . (I) Transition topology for “deterministic cyclical process”;
. (I) Transition topology for “deterministic cyclical process”;  . The transition topologies in (G) to (I) were refuted given the strong support for topologies in (D) and (F) against their rival hypotheses. For parameter values see Table S3.
. The transition topologies in (G) to (I) were refuted given the strong support for topologies in (D) and (F) against their rival hypotheses. For parameter values see Table S3.
 . (B) Distribution for molecules isolated from pho4[85-99] pho80Δ cells with a mutated PHO5 TATA box. PHO5 expression in pho4[85-99] pho80Δ cells with wild type PHO5 TATA is 0.14 relative to PHO4 wild type (set to 1) . Predictions were based on the topology of Figure 4F, rather than 4D;
. (B) Distribution for molecules isolated from pho4[85-99] pho80Δ cells with a mutated PHO5 TATA box. PHO5 expression in pho4[85-99] pho80Δ cells with wild type PHO5 TATA is 0.14 relative to PHO4 wild type (set to 1) . Predictions were based on the topology of Figure 4F, rather than 4D;  . (C) Distribution for molecules isolated from pho4Δ pho80Δ cells with a mutated PHO5 TATA box. Expression of PHO5 in pho4Δ pho80Δ cells relative to PHO4 wild type is ∼0.005 (see below and [15]). Predictions were based on the topology of Figure 4D, rather than 4F;
. (C) Distribution for molecules isolated from pho4Δ pho80Δ cells with a mutated PHO5 TATA box. Expression of PHO5 in pho4Δ pho80Δ cells relative to PHO4 wild type is ∼0.005 (see below and [15]). Predictions were based on the topology of Figure 4D, rather than 4F;  . (D) Distribution for molecules isolated from PHO4 pho2Δ cells grown in high phosphate (PHO4cyt); Pho4 is in the cytoplasm, rather than the nucleus, and PHO5 is repressed, therefore. Predictions were based on the topology of Figure 4D, rather than 4F;
. (D) Distribution for molecules isolated from PHO4 pho2Δ cells grown in high phosphate (PHO4cyt); Pho4 is in the cytoplasm, rather than the nucleus, and PHO5 is repressed, therefore. Predictions were based on the topology of Figure 4D, rather than 4F;  . (E) Distribution for molecules isolated from PHO4 cells grown in high phosphate (PHO4cyt). Predictions were based on the topology of Figure 4D, rather than 4F;
. (E) Distribution for molecules isolated from PHO4 cells grown in high phosphate (PHO4cyt). Predictions were based on the topology of Figure 4D, rather than 4F;  . For parameter values see Table S3.
. For parameter values see Table S3.
 min−1,
min−1,  h−1,
h−1,  h−1 (see Figure 1A, and main text below; like Pho5, CFP and YFP are biochemically stable; the proteins are lost therefore primarily due to dilution by cell division). With
h−1 (see Figure 1A, and main text below; like Pho5, CFP and YFP are biochemically stable; the proteins are lost therefore primarily due to dilution by cell division). With  , the kinetic parameter for transcription is
, the kinetic parameter for transcription is  min−1. The parameter values were determined as described in the main text.
min−1. The parameter values were determined as described in the main text.
 h−1. A closely similar value for δ was obtained for the PHO5 mRNA (data not shown). The steady state abundance of PHO5 transcripts is therefore expected to be similar to the number of CFP mRNAs per cell.
h−1. A closely similar value for δ was obtained for the PHO5 mRNA (data not shown). The steady state abundance of PHO5 transcripts is therefore expected to be similar to the number of CFP mRNAs per cell.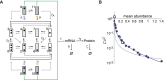
 ,
,  , and λ as regulatory parameters, which were allowed to float along the line
, and λ as regulatory parameters, which were allowed to float along the line  , with
, with  ,
,  , where the hat and prime mark the parameter values for the PHO4 wild type and pho4Δ mutant, respectively, and t is a real number ≥0. Noise predictions based on the assumption of
, where the hat and prime mark the parameter values for the PHO4 wild type and pho4Δ mutant, respectively, and t is a real number ≥0. Noise predictions based on the assumption of  ,
,  , and ε as regulatory parameters, and thus assuming a combination of burst size and burst frequency control, are indicated by the gray curve. Virtually the same result was obtained on the topology of Figure 4F for nucleosome transition (not shown).
, and ε as regulatory parameters, and thus assuming a combination of burst size and burst frequency control, are indicated by the gray curve. Virtually the same result was obtained on the topology of Figure 4F for nucleosome transition (not shown).Comment in
-
Bursting with randomness: a simple model for stochastic control of gene expression.PLoS Biol. 2013;11(8):e1001622. doi: 10.1371/journal.pbio.1001622. Epub 2013 Aug 6. PLoS Biol. 2013. PMID: 23940459 Free PMC article. No abstract available.
References
-
- Elowitz MB, Levine AJ, Siggia ED, Swain PS (2002) Stochastic gene expression in a single cell. Science 297: 1183–1186. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

