A LacI-family regulator activates maltodextrin metabolism of Enterococcus faecium
- PMID: 23951303
- PMCID: PMC3737153
- DOI: 10.1371/journal.pone.0072285
A LacI-family regulator activates maltodextrin metabolism of Enterococcus faecium
Abstract
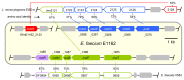
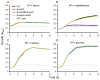
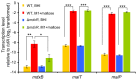
Enterococcus faecium is a gut commensal of humans and animals. In the intestinal tract, E. faecium will have access to a wide variety of carbohydrates, including maltodextrins and maltose, which are the sugars that result from the enzymatic digestion of starch by host-derived and microbial amylases. In this study, we identified the genetic determinants for maltodextrin utilization of E. faecium E1162. We generated a deletion mutant of the mdxABCD-pulA gene cluster that is homologous to maltodextrin uptake genes in other Gram-positive bacteria, and a deletion mutant of the mdxR gene, which is predicted to encode a LacI family regulator of mdxABCD-pulA. Both mutations impaired growth on maltodextrins but had no effect on the growth on maltose and glucose. Comparative transcriptome analysis showed that eight genes (including mdxABCD-pulA) were expressed at significantly lower levels in the isogenic ΔmdxR mutant strain compared to the parental strain when grown on maltose. Quantitative real-time RT-PCR confirmed the results of transcriptome analysis and showed that the transcription of a putative maltose utilization gene cluster is induced in a semi-defined medium supplemented with maltose but is not regulated by MdxR. Understanding the maltodextrin metabolism of E. faecium could yield novel insights into the underlying mechanisms that contribute to the gut commensal lifestyle of E. faecium.
Conflict of interest statement
Figures
Similar articles
-
Enterococcus faecalis Maltodextrin Gene Regulation by Combined Action of Maltose Gene Regulator MalR and Pleiotropic Regulator CcpA.Appl Environ Microbiol. 2020 Sep 1;86(18):e01147-20. doi: 10.1128/AEM.01147-20. Print 2020 Sep 1. Appl Environ Microbiol. 2020. PMID: 32680872 Free PMC article.
-
The N-terminal domain of the thermo-regulated surface protein PrpA of Enterococcus faecium binds to fibrinogen, fibronectin and platelets.Sci Rep. 2015 Dec 17;5:18255. doi: 10.1038/srep18255. Sci Rep. 2015. PMID: 26675410 Free PMC article.
-
Enterococcus faecalis Uses a Phosphotransferase System Permease and a Host Colonization-Related ABC Transporter for Maltodextrin Uptake.J Bacteriol. 2017 Apr 11;199(9):e00878-16. doi: 10.1128/JB.00878-16. Print 2017 May 1. J Bacteriol. 2017. PMID: 28242718 Free PMC article.
-
Maltose and maltodextrin utilization by Listeria monocytogenes depend on an inducible ABC transporter which is repressed by glucose.PLoS One. 2010 Apr 27;5(4):e10349. doi: 10.1371/journal.pone.0010349. PLoS One. 2010. PMID: 20436965 Free PMC article.
-
Functional genomic analysis of bile salt resistance in Enterococcus faecium.BMC Genomics. 2013 May 3;14:299. doi: 10.1186/1471-2164-14-299. BMC Genomics. 2013. PMID: 23641968 Free PMC article.
Cited by
-
Enterococcus faecalis Maltodextrin Gene Regulation by Combined Action of Maltose Gene Regulator MalR and Pleiotropic Regulator CcpA.Appl Environ Microbiol. 2020 Sep 1;86(18):e01147-20. doi: 10.1128/AEM.01147-20. Print 2020 Sep 1. Appl Environ Microbiol. 2020. PMID: 32680872 Free PMC article.
-
The N-terminal domain of the thermo-regulated surface protein PrpA of Enterococcus faecium binds to fibrinogen, fibronectin and platelets.Sci Rep. 2015 Dec 17;5:18255. doi: 10.1038/srep18255. Sci Rep. 2015. PMID: 26675410 Free PMC article.
-
The Fe-S cluster biosynthesis in Enterococcus faecium is essential for anaerobic growth and gastrointestinal colonization.Gut Microbes. 2024 Jan-Dec;16(1):2359665. doi: 10.1080/19490976.2024.2359665. Epub 2024 Jun 3. Gut Microbes. 2024. PMID: 38831611 Free PMC article.
-
Enterococcus faecalis Uses a Phosphotransferase System Permease and a Host Colonization-Related ABC Transporter for Maltodextrin Uptake.J Bacteriol. 2017 Apr 11;199(9):e00878-16. doi: 10.1128/JB.00878-16. Print 2017 May 1. J Bacteriol. 2017. PMID: 28242718 Free PMC article.
-
Plasmids encode niche-specific traits in Lactobacillaceae.Microb Genom. 2021 Mar;7(3):mgen000472. doi: 10.1099/mgen.0.000472. Epub 2020 Nov 9. Microb Genom. 2021. PMID: 33166245 Free PMC article.
References
-
- Top J, Willems R, Bonten M (2008) Emergence of CC17 Enterococcus faecium: from commensal to hospital-adapted pathogen. FEMS Immunol Med Microbiol 52: 297-308. doi:10.1111/j.1348-0421.2008.00039.x. PubMed: 18279340. - DOI - PubMed
-
- Willems RJ, van Schaik W (2009) Transition of Enterococcus faecium from commensal organism to nosocomial pathogen. Future Microbiol 4: 1125-1135. doi:10.2217/fmb.09.82. PubMed: 19895216. - DOI - PubMed
-
- Chang DE, Smalley DJ, Tucker DL, Leatham MP, Norris WE et al. (2004) Carbon nutrition of Escherichia coli in the mouse intestine. Proc Natl Acad Sci U S A 101: 7427-7432. doi:10.1073/pnas.0307888101. PubMed: 15123798. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

