Precise developmental gene expression arises from globally stochastic transcriptional activity
- PMID: 23953111
- PMCID: PMC3778922
- DOI: 10.1016/j.cell.2013.07.025
Precise developmental gene expression arises from globally stochastic transcriptional activity
Abstract
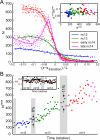
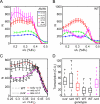
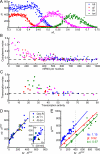
Early embryonic patterning events are strikingly precise, a fact that appears incompatible with the stochastic gene expression observed across phyla. Using single-molecule mRNA quantification in Drosophila embryos, we determine the magnitude of fluctuations in the expression of four critical patterning genes. The accumulation of mRNAs is identical across genes and fluctuates by only ∼8% between neighboring nuclei, generating precise protein distributions. In contrast, transcribing loci exhibit an intrinsic noise of ∼45% independent of specific promoter-enhancer architecture or fluctuating inputs. Precise transcript distribution in the syncytium is recovered via straightforward spatiotemporal averaging, i.e., accumulation and diffusion of transcripts during nuclear cycles, without regulatory feedback. Common expression characteristics shared between genes suggest that fluctuations in mRNA production are context independent and are a fundamental property of transcription. The findings shed light on how the apparent paradox between stochastic transcription and developmental precision is resolved.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures



References
-
- Arias AM, Hayward P. Filtering transcriptional noise during development: concepts and mechanisms. Nat Rev Genet. 2006;7:34–44. - PubMed
-
- Balaban NQ, Merrin J, Chait R, Kowalik L, Leibler S. Bacterial persistence as a phenotypic switch. Science. 2004;305:1622–1625. - PubMed
-
- Bar-Even A, Paulsson J, Maheshri N, Carmi M, O'Shea E, Pilpel Y, Barkai N. Noise in protein expression scales with natural protein abundance. Nat Genet. 2006;38:636–643. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

