Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria
- PMID: 23954787
- PMCID: PMC3770465
- DOI: 10.1016/j.celrep.2013.07.026
Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria
Abstract
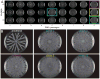
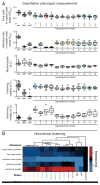
Most bacteria in nature live in surface-associated communities rather than planktonic populations. Nonetheless, how surface-associated environments shape bacterial evolutionary adaptation remains poorly understood. Here, we show that subjecting Pseudomonas aeruginosa to repeated rounds of swarming, a collective form of surface migration, drives remarkable parallel evolution toward a hyperswarmer phenotype. In all independently evolved hyperswarmers, the reproducible hyperswarming phenotype is caused by parallel point mutations in a flagellar synthesis regulator, FleN, which locks the naturally monoflagellated bacteria in a multiflagellated state and confers a growth rate-independent advantage in swarming. Although hyperswarmers outcompete the ancestral strain in swarming competitions, they are strongly outcompeted in biofilm formation, which is an essential trait for P. aeruginosa in environmental and clinical settings. The finding that evolution in swarming colonies reliably produces evolution of poor biofilm formers supports the existence of an evolutionary trade-off between motility and biofilm formation.
Copyright © 2013 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Comment in
-
You get what you select for: better swarming through more flagella.Trends Microbiol. 2013 Oct;21(10):508-9. doi: 10.1016/j.tim.2013.08.003. Epub 2013 Sep 16. Trends Microbiol. 2013. PMID: 24051005
Similar articles
-
Hyperswarming adaptations in a bacterium improve collective motility without enhancing single cell motility.Soft Matter. 2014 Apr 14;10(14):2405-13. doi: 10.1039/c3sm53127a. Soft Matter. 2014. PMID: 24622509 Free PMC article.
-
Swarming of Pseudomonas aeruginosa is controlled by a broad spectrum of transcriptional regulators, including MetR.J Bacteriol. 2009 Sep;191(18):5592-602. doi: 10.1128/JB.00157-09. Epub 2009 Jul 10. J Bacteriol. 2009. PMID: 19592586 Free PMC article.
-
The evolutionary trajectories of P. aeruginosa in biofilm and planktonic growth modes exposed to ciprofloxacin: beyond selection of antibiotic resistance.NPJ Biofilms Microbiomes. 2020 Jul 24;6(1):28. doi: 10.1038/s41522-020-00138-8. NPJ Biofilms Microbiomes. 2020. PMID: 32709907 Free PMC article.
-
Rapid evolution of culture-impaired bacteria during adaptation to biofilm growth.Cell Rep. 2014 Jan 30;6(2):293-300. doi: 10.1016/j.celrep.2013.12.019. Epub 2014 Jan 9. Cell Rep. 2014. PMID: 24412364 Free PMC article.
-
The Ultimate Guide to Bacterial Swarming: An Experimental Model to Study the Evolution of Cooperative Behavior.Annu Rev Microbiol. 2019 Sep 8;73:293-312. doi: 10.1146/annurev-micro-020518-120033. Epub 2019 Jun 10. Annu Rev Microbiol. 2019. PMID: 31180806 Free PMC article. Review.
Cited by
-
Transformed Recombinant Enrichment Profiling Rapidly Identifies HMW1 as an Intracellular Invasion Locus in Haemophilus influenza.PLoS Pathog. 2016 Apr 28;12(4):e1005576. doi: 10.1371/journal.ppat.1005576. eCollection 2016 Apr. PLoS Pathog. 2016. PMID: 27124727 Free PMC article.
-
Insights into the Oxidative Stress Response of Salmonella enterica serovar Enteritidis Revealed by the Next Generation Sequencing Approach.Antioxidants (Basel). 2020 Sep 10;9(9):849. doi: 10.3390/antiox9090849. Antioxidants (Basel). 2020. PMID: 32927804 Free PMC article.
-
Experimental evolution reveals nitrate tolerance mechanisms in Desulfovibrio vulgaris.ISME J. 2020 Nov;14(11):2862-2876. doi: 10.1038/s41396-020-00753-5. Epub 2020 Sep 15. ISME J. 2020. PMID: 32934357 Free PMC article.
-
Cell-Size Homeostasis and the Incremental Rule in a Bacterial Pathogen.Biophys J. 2015 Aug 4;109(3):521-8. doi: 10.1016/j.bpj.2015.07.002. Biophys J. 2015. PMID: 26244734 Free PMC article.
-
Learning the space-time phase diagram of bacterial swarm expansion.Proc Natl Acad Sci U S A. 2019 Jan 29;116(5):1489-1494. doi: 10.1073/pnas.1811722116. Epub 2019 Jan 11. Proc Natl Acad Sci U S A. 2019. PMID: 30635422 Free PMC article.
References
-
- Boles BR, Thoendel M, Singh PK. Rhamnolipids mediate detachment of Pseudomonas aeruginosa from biofilms. Molecular microbiology. 2005;57:1210–1223. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

