E2 ubiquitin-conjugating enzymes regulate the deubiquitinating activity of OTUB1
- PMID: 23955022
- PMCID: PMC3941643
- DOI: 10.1038/nsmb.2655
E2 ubiquitin-conjugating enzymes regulate the deubiquitinating activity of OTUB1
Abstract
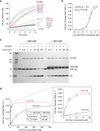
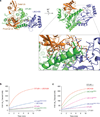
OTUB1 is a Lys48-specific deubiquitinating enzyme that forms a complex in vivo with E2 ubiquitin (Ub)-conjugating enzymes including UBC13 and UBCH5. OTUB1 binds E2~Ub thioester intermediates and prevents ubiquitin transfer, thereby noncatalytically inhibiting accumulation of polyubiquitin. We report here that a second role of OTUB1-E2 interactions is to stimulate OTUB1 cleavage of Lys48 polyubiquitin. This stimulation is regulated by the ratio of charged to uncharged E2 and by the concentration of Lys48-linked polyubiquitin and free ubiquitin. Structural and biochemical studies of human and worm OTUB1 and UBCH5B show that the E2 enzyme stimulates binding of the Lys48 polyubiquitin substrate by stabilizing folding of the OTUB1 N-terminal ubiquitin-binding helix. Our results suggest that OTUB1-E2 complexes in the cell are poised to regulate polyubiquitin chain elongation or degradation in response to changing levels of E2 charging and available free ubiquitin.
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

