A genomic-scale artificial microRNA library as a tool to investigate the functionally redundant gene space in Arabidopsis
- PMID: 23956262
- PMCID: PMC3784584
- DOI: 10.1105/tpc.113.112805
A genomic-scale artificial microRNA library as a tool to investigate the functionally redundant gene space in Arabidopsis
Abstract
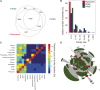
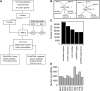
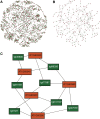
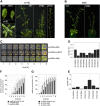
Traditional forward genetic screens are limited in the identification of homologous genes with overlapping functions. Here, we report the analyses and assembly of genome-wide protein family definitions that comprise the largest estimate for the potentially redundant gene space in Arabidopsis thaliana. On this basis, a computational design of genome-wide family-specific artificial microRNAs (amiRNAs) was performed using high-performance computing resources. The amiRNA designs are searchable online (http://phantomdb.ucsd.edu). A computationally derived library of 22,000 amiRNAs was synthesized in 10 sublibraries of 1505 to 4082 amiRNAs, each targeting defined functional protein classes. For example, 2964 amiRNAs target annotated DNA and RNA binding protein families and 1777 target transporter proteins, and another sublibrary targets proteins of unknown function. To evaluate the potential of an amiRNA-based screen, we tested 122 amiRNAs targeting transcription factor, protein kinase, and protein phosphatase families. Several amiRNA lines showed morphological phenotypes, either comparable to known phenotypes of single and double/triple mutants or caused by overexpression of microRNAs. Moreover, novel morphological and abscisic acid-insensitive seed germination mutants were identified for amiRNAs targeting zinc finger homeodomain transcription factors and mitogen-activated protein kinase kinase kinases, respectively. These resources provide an approach for genome-wide genetic screens of the functionally redundant gene space in Arabidopsis.
Figures

References
-
- Abe M., Katsumata H., Komeda Y., Takahashi T. (2003). Regulation of shoot epidermal cell differentiation by a pair of homeodomain proteins in Arabidopsis. Development 130: 635–643 - PubMed
-
- Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. (1990). Basic local alignment search tool. J. Mol. Biol. 215: 403–410 - PubMed
-
- Arabidopsis Genome Initiative (2000). Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

