The N-degradome of Escherichia coli: limited proteolysis in vivo generates a large pool of proteins bearing N-degrons
- PMID: 23960079
- PMCID: PMC3789986
- DOI: 10.1074/jbc.M113.492108
The N-degradome of Escherichia coli: limited proteolysis in vivo generates a large pool of proteins bearing N-degrons
Abstract
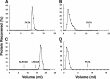
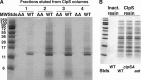
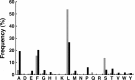
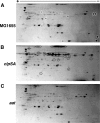
The N-end rule is a conserved mechanism found in Gram-negative bacteria and eukaryotes for marking proteins to be degraded by ATP-dependent proteases. Specific N-terminal amino acids (N-degrons) are sufficient to target a protein to the degradation machinery. In Escherichia coli, the adaptor ClpS binds an N-degron and delivers the protein to ClpAP for degradation. As ClpS recognizes N-terminal Phe, Trp, Tyr, and Leu, which are not found at the N terminus of proteins translated and processed by the canonical pathway, proteins must be post-translationally modified to expose an N-degron. One modification is catalyzed by Aat, an enzyme that adds leucine or phenylalanine to proteins with N-terminal lysine or arginine; however, such proteins are also not generated by the canonical protein synthesis pathway. Thus, the mechanisms producing N-degrons in proteins and the frequency of their occurrence largely remain a mystery. To address these issues, we used a ClpS affinity column to isolate interacting proteins from E. coli cell lysates under non-denaturing conditions. We identified more than 100 proteins that differentially bound to a column charged with wild-type ClpS and eluted with a peptide bearing an N-degron. Thirty-two of 37 determined N-terminal peptides had N-degrons. Most of the proteins were N-terminally truncated by endoproteases or exopeptidases, and many were further modified by Aat. The identities of the proteins point to possible physiological roles for the N-end rule in cell division, translation, transcription, and DNA replication and reveal widespread proteolytic processing of cellular proteins to generate N-end rule substrates.
Keywords: ATP-dependent Protease; Aat; Adaptor Proteins; ClpA; ClpS; END Site; N-end Rule; Protein Degradation; Protein Processing; Proteomics.
Figures
Similar articles
-
Structural basis of N-end rule substrate recognition in Escherichia coli by the ClpAP adaptor protein ClpS.EMBO Rep. 2009 May;10(5):508-14. doi: 10.1038/embor.2009.62. Epub 2009 Apr 17. EMBO Rep. 2009. PMID: 19373253 Free PMC article.
-
ClpS is an essential component of the N-end rule pathway in Escherichia coli.Nature. 2006 Feb 9;439(7077):753-6. doi: 10.1038/nature04412. Nature. 2006. PMID: 16467841
-
Modification of PATase by L/F-transferase generates a ClpS-dependent N-end rule substrate in Escherichia coli.EMBO J. 2009 Jun 17;28(12):1732-44. doi: 10.1038/emboj.2009.134. Epub 2009 May 14. EMBO J. 2009. PMID: 19440203 Free PMC article.
-
N-degron and C-degron pathways of protein degradation.Proc Natl Acad Sci U S A. 2019 Jan 8;116(2):358-366. doi: 10.1073/pnas.1816596116. Proc Natl Acad Sci U S A. 2019. PMID: 30622213 Free PMC article. Review.
-
The N-end rule pathway and regulation by proteolysis.Protein Sci. 2011 Aug;20(8):1298-345. doi: 10.1002/pro.666. Protein Sci. 2011. PMID: 21633985 Free PMC article. Review.
Cited by
-
Kinetics of Bovine leukemia virus aspartic protease reveals its dimerization and conformational change.PLoS One. 2022 Jul 22;17(7):e0271671. doi: 10.1371/journal.pone.0271671. eCollection 2022. PLoS One. 2022. PMID: 35867649 Free PMC article.
-
Perspectives and Insights into the Competition for Aminoacyl-tRNAs between the Translational Machinery and for tRNA Dependent Non-Ribosomal Peptide Bond Formation.Life (Basel). 2015 Dec 31;6(1):2. doi: 10.3390/life6010002. Life (Basel). 2015. PMID: 26729173 Free PMC article. Review.
-
Structural Basis of an N-Degron Adaptor with More Stringent Specificity.Structure. 2016 Feb 2;24(2):232-42. doi: 10.1016/j.str.2015.12.008. Epub 2016 Jan 21. Structure. 2016. PMID: 26805523 Free PMC article.
-
Degradation of Serotonin N-Acetyltransferase, a Circadian Regulator, by the N-end Rule Pathway.J Biol Chem. 2016 Aug 12;291(33):17178-96. doi: 10.1074/jbc.M116.734640. Epub 2016 Jun 23. J Biol Chem. 2016. PMID: 27339900 Free PMC article.
-
The lowdown on breakdown: Open questions in plant proteolysis.Plant Cell. 2024 Sep 3;36(9):2931-2975. doi: 10.1093/plcell/koae193. Plant Cell. 2024. PMID: 38980154 Free PMC article. Review.
References
-
- Gottesman S., Wickner S., Maurizi M. R. (1997) Protein quality control: triage by chaperones and proteases. Genes Dev. 11, 815–823 - PubMed
-
- Dougan D. A., Mogk A., Zeth K., Turgay K., Bukau B. (2002) AAA+ proteins and substrate recognition, it all depends on their partner in crime. FEBS Lett. 529, 6–10 - PubMed
-
- Kessel M., Maurizi M. R., Kim B., Kocsis E., Trus B. L., Singh S. K., Steven A. C. (1995) Homology in structural organization between E. coli ClpAP protease and the eukaryotic 26 S proteasome. J. Mol. Biol. 250, 587–594 - PubMed
-
- Wang J., Hartling J. A., Flanagan J. M. (1997) The structure of ClpP at 2.3 Å resolution suggests a model for ATP-dependent proteolysis. Cell 91, 447–456 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

