Bioinformatic analyses of integral membrane transport proteins encoded within the genome of the planctomycetes species, Rhodopirellula baltica
- PMID: 23969110
- PMCID: PMC3905805
- DOI: 10.1016/j.bbamem.2013.08.007
Bioinformatic analyses of integral membrane transport proteins encoded within the genome of the planctomycetes species, Rhodopirellula baltica
Abstract
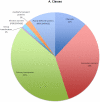
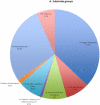
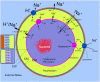
Rhodopirellula baltica (R. baltica) is a Planctomycete, known to have intracellular membranes. Because of its unusual cell structure and ecological significance, we have conducted comprehensive analyses of its transmembrane transport proteins. The complete proteome of R. baltica was screened against the Transporter Classification Database (TCDB) to identify recognizable integral membrane transport proteins. 342 proteins were identified with a high degree of confidence, and these fell into several different classes. R. baltica encodes in its genome channels (12%), secondary carriers (33%), and primary active transport proteins (41%) in addition to classes represented in smaller numbers. Relative to most non-marine bacteria, R. baltica possesses a larger number of sodium-dependent symporters but fewer proton-dependent symporters, and it has dimethylsulfoxide (DMSO) and trimethyl-amine-oxide (TMAO) reductases, consistent with its Na(+)-rich marine environment. R. baltica also possesses a Na(+)-translocating NADH:quinone dehydrogenase (Na(+)-NDH), a Na(+) efflux decarboxylase, two Na(+)-exporting ABC pumps, two Na(+)-translocating F-type ATPases, two Na(+):H(+) antiporters and two K(+):H(+) antiporters. Flagellar motility probably depends on the sodium electrochemical gradient. Surprisingly, R. baltica also has a complete set of H(+)-translocating electron transport complexes similar to those present in α-proteobacteria and eukaryotic mitochondria. The transport proteins identified proved to be typical of the bacterial domain with little or no indication of the presence of eukaryotic-type transporters. However, novel functionally uncharacterized multispanning membrane proteins were identified, some of which are found only in Rhodopirellula species, but others of which are widely distributed in bacteria. The analyses lead to predictions regarding the physiology, ecology and evolution of R. baltica.
Keywords: Cellular energization; Electron transport; Marine ecology; Planctomycetes; Sodium motive force; Transport proteins.
© 2013.
Figures


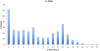
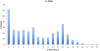
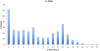
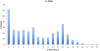
References
-
- Rabus R. An overview of 2D DIGE analysis of marine (environmental) bacteria. Methods Mol Biol. 2012;854:355–372. - PubMed
-
- Lindsay MR, Webb RI, Strous M, Jetten MS, Butler MK, Forde RJ, Fuerst JA. Cell compartmentalisation in planctomycetes: novel types of structural organisation for the bacterial cell. Arch Microbiol. 2001;175:413–429. - PubMed
-
- Lindsay MR, Webb RI, Fuerst JA. Pirellulosomes: a new type of membrane-bounded cell compartment in planctomycete bacteria of the genus Pirellula. Microbiology. 1997;143:739–748. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

