Fungi of the murine gut: episodic variation and proliferation during antibiotic treatment
- PMID: 23977147
- PMCID: PMC3747063
- DOI: 10.1371/journal.pone.0071806
Fungi of the murine gut: episodic variation and proliferation during antibiotic treatment
Abstract
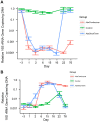
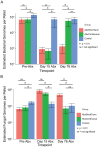
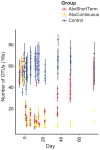
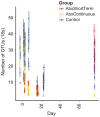
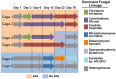
Antibiotic use in humans has been associated with outgrowth of fungi. Here we used a murine model to investigate the gut microbiome over 76 days of treatment with vancomycin, ampicillin, neomycin, and metronidazole and subsequent recovery. Mouse stool was studied as a surrogate for the microbiota of the lower gastrointestinal tract. The abundance of fungi and bacteria was measured using quantitative PCR, and the proportional composition of the communities quantified using 454/Roche pyrosequencing of rRNA gene tags. Prior to treatment, bacteria outnumbered fungi by >3 orders of magnitude. Upon antibiotic treatment, bacteria dropped in abundance >3 orders of magnitude, so that the predominant 16S sequences detected became transients derived from food. Upon cessation of treatment, bacterial communities mostly returned to their previous numbers and types after 8 weeks, though communities remained detectably different from untreated controls. Fungal communities varied substantially over time, even in the untreated controls. Separate cages within the same treatment group showed radical differences, but mice within a cage generally behaved similarly. Fungi increased ∼40-fold in abundance upon antibiotic treatment but declined back to their original abundance after cessation of treatment. At the last time point, Candida remained more abundant than prior to treatment. These data show that 1) gut fungal populations change radically during normal mouse husbandry, 2) fungi grow out in the gut upon suppression of bacterial communities with antibiotics, and 3) perturbations due to antibiotics persist long term in both the fungal and bacterial microbiota.
Conflict of interest statement
Figures



References
-
- Chen KY, Ko SC, Hsueh PR, Luh KT, Yang PC (2001) Pulmonary fungal infection: emphasis on microbiological spectra, patient outcome, and prognostic factors. Chest 120: 177–184. - PubMed
-
- Abbott J (1995) Clinical and microscopic diagnosis of vaginal yeast infection: a prospective analysis. Annals of emergency medicine 25: 587–591. - PubMed
-
- Oksala E (1990) Factors predisposing to oral yeast infections. Acta odontologica Scandinavica 48: 71–74. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

