DNA double strand break repair via non-homologous end-joining
- PMID: 24000320
- PMCID: PMC3758668
- DOI: 10.3978/j.issn.2218-676X.2013.04.02
DNA double strand break repair via non-homologous end-joining
Abstract
DNA double-stranded breaks (DSB) are among the most dangerous forms of DNA damage. Unrepaired DSBs results in cells undergoing apoptosis or senescence whereas mis-processing of DSBs can lead to genomic instability and carcinogenesis. One important pathway in eukaryotic cells responsible for the repair of DSBs is non-homologous end-joining (NHEJ). In this review we will discuss the interesting new insights into the mechanism of the NHEJ pathway and the proteins which mediate this repair process. Furthermore, the general role of NHEJ in promoting genomic stability will be discussed.
Keywords: DNA double strand breaks; DNA-Ligase IV; DNA-PKcs; Ku70/80; XLF; XRCC4; non-homologous end-joining.
Conflict of interest statement
Figures

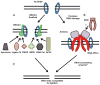
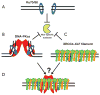
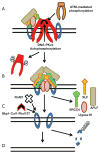
References
-
- Khanna KK, Jackson SP. DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet. 2001;27:247–54. - PubMed
-
- Hoeijmakers JH. Genome maintenance mechanisms for preventing cancer. Nature. 2001;411:366–74. - PubMed
-
- Malu S, Malshetty V, Francis D, et al. Role of non-homologous end joining in V(D)J recombination. Immunol Res. 2012;54:233–46. - PubMed
-
- Burma S, Chen BP, Chen DJ. Role of non-homologous end joining (NHEJ) in maintaining genomic integrity. DNA Repair (Amst) 2006;5:1042–8. - PubMed
-
- Weterings E, Chen DJ. The endless tale of non-homologous end-joining. Cell Res. 2008;18:114–24. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
