Ecology, diversity, and evolution of magnetotactic bacteria
- PMID: 24006473
- PMCID: PMC3811606
- DOI: 10.1128/MMBR.00021-13
Ecology, diversity, and evolution of magnetotactic bacteria
Abstract
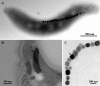
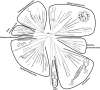
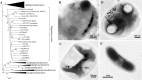
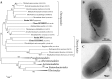
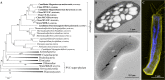
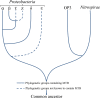
Magnetotactic bacteria (MTB) are widespread, motile, diverse prokaryotes that biomineralize a unique organelle called the magnetosome. Magnetosomes consist of a nano-sized crystal of a magnetic iron mineral that is enveloped by a lipid bilayer membrane. In cells of almost all MTB, magnetosomes are organized as a well-ordered chain. The magnetosome chain causes the cell to behave like a motile, miniature compass needle where the cell aligns and swims parallel to magnetic field lines. MTB are found in almost all types of aquatic environments, where they can account for an important part of the bacterial biomass. The genes responsible for magnetosome biomineralization are organized as clusters in the genomes of MTB, in some as a magnetosome genomic island. The functions of a number of magnetosome genes and their associated proteins in magnetosome synthesis and construction of the magnetosome chain have now been elucidated. The origin of magnetotaxis appears to be monophyletic; that is, it developed in a common ancestor to all MTB, although horizontal gene transfer of magnetosome genes also appears to play a role in their distribution. The purpose of this review, based on recent progress in this field, is focused on the diversity and the ecology of the MTB and also the evolution and transfer of the molecular determinants involved in magnetosome formation.
Figures
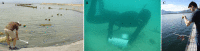






Similar articles
-
The bacterial magnetosome: a unique prokaryotic organelle.J Mol Microbiol Biotechnol. 2013;23(1-2):63-80. doi: 10.1159/000346543. Epub 2013 Apr 18. J Mol Microbiol Biotechnol. 2013. PMID: 23615196 Review.
-
Expanding magnetic organelle biogenesis in the domain Bacteria.Microbiome. 2020 Oct 30;8(1):152. doi: 10.1186/s40168-020-00931-9. Microbiome. 2020. PMID: 33126926 Free PMC article.
-
Monophyletic origin of magnetotaxis and the first magnetosomes.Environ Microbiol. 2013 Aug;15(8):2267-74. doi: 10.1111/1462-2920.12097. Epub 2013 Feb 25. Environ Microbiol. 2013. PMID: 23438345
-
Morphological and cellular diversity of magnetotactic bacteria: A review.J Basic Microbiol. 2018 May;58(5):378-389. doi: 10.1002/jobm.201700383. Epub 2017 Nov 7. J Basic Microbiol. 2018. PMID: 29112284 Review.
-
Common ancestry of iron oxide- and iron-sulfide-based biomineralization in magnetotactic bacteria.ISME J. 2011 Oct;5(10):1634-40. doi: 10.1038/ismej.2011.35. Epub 2011 Apr 21. ISME J. 2011. PMID: 21509043 Free PMC article.
Cited by
-
Ferromagnetic resonance spectra of linear magnetosome chains.Beilstein J Nanotechnol. 2024 Feb 5;15:157-167. doi: 10.3762/bjnano.15.15. eCollection 2024. Beilstein J Nanotechnol. 2024. PMID: 38352719 Free PMC article.
-
Preliminary indication of the role of AHL-dependent quorum sensing systems in calcium carbonate precipitation in Gram-negative bacteria.AIMS Microbiol. 2023 Sep 26;9(4):692-711. doi: 10.3934/microbiol.2023035. eCollection 2023. AIMS Microbiol. 2023. PMID: 38173968 Free PMC article.
-
Positioning the flagellum at the center of a dividing cell to combine bacterial division with magnetic polarity.mBio. 2015 Feb 24;6(2):e02286. doi: 10.1128/mBio.02286-14. mBio. 2015. PMID: 25714711 Free PMC article.
-
Magnetic Entropy as a Proposed Gating Mechanism for Magnetogenetic Ion Channels.Biophys J. 2019 Feb 5;116(3):454-468. doi: 10.1016/j.bpj.2019.01.003. Epub 2019 Jan 8. Biophys J. 2019. PMID: 30665695 Free PMC article.
-
Biogeochemical Niche of Magnetotactic Cocci Capable of Sequestering Large Polyphosphate Inclusions in the Anoxic Layer of the Lake Pavin Water Column.Front Microbiol. 2022 Jan 10;12:789134. doi: 10.3389/fmicb.2021.789134. eCollection 2021. Front Microbiol. 2022. PMID: 35082768 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

