Using ion mobility spectrometry-mass spectrometry to decipher the conformational and assembly characteristics of the hepatitis B capsid protein
- PMID: 24010669
- PMCID: PMC3762345
- DOI: 10.1016/j.bpj.2013.07.028
Using ion mobility spectrometry-mass spectrometry to decipher the conformational and assembly characteristics of the hepatitis B capsid protein
Abstract
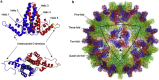
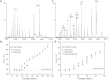
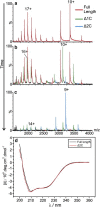
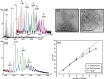
The structural and functional analysis of the core protein of hepatitis B virus is important for a full understanding of the viral life cycle and the development of novel therapeutic agents. The majority of the core protein (CP149) comprises the capsid assembly domain, and the C-terminal region (residues 150-183) is responsible for nucleic acid binding. Protein monomers associate to form dimeric structural subunits, and helices 3 and 4 (residues 50-111 of the assembly domain) have been shown to be important for this as they constitute the interdimer interface. Here, using mass spectrometry coupled with ion mobility spectrometry, we demonstrate the conformational flexibility of the CP149 dimer. Limited proteolysis was used to locate involvement in this feature to the C-terminal region. A genetically fused CP dimer was found to show decreased disorder, consistent with a more restricted C-terminus at the fusion junction. Incubation of CP149 dimer with heteroaryldihydropyrimidine-1, a small molecule known to interfere with the assembly process, was shown to result in oligomers different in shape to the capsid assembly-competent oligomers of the fused CP dimer. We suggest that heteroaryldihydropyrimidine-1 affects the dynamics of CP149 dimer in solution, likely affecting the ratio between assembly active and inactive states. Therefore, assembly of the less dynamic fused dimer is less readily misdirected by heteroaryldihydropyrimidine-1. These studies of the flexibility and oligomerization properties of hepatitis B virus core protein illustrate both the importance of C-terminal dynamics in function and the utility of gas-phase techniques for structural and dynamical biomolecular analysis.
Copyright © 2013 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures

References
-
- Dienstag J.L. Hepatitis B virus infection. N. Engl. J. Med. 2008;359:1486–1500. - PubMed
-
- Nassal M., Schaller H. Hepatitis B virus replication. Trends Microbiol. 1993;1:221–228. - PubMed
-
- Stray S.J., Ceres P., Zlotnick A. Zinc ions trigger conformational change and oligomerization of hepatitis B virus capsid protein. Biochemistry. 2004;43:9989–9998. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

