The COMBREX project: design, methodology, and initial results
- PMID: 24013487
- PMCID: PMC3754883
- DOI: 10.1371/journal.pbio.1001638
The COMBREX project: design, methodology, and initial results
Abstract
Experimental data exists for only a vanishingly small fraction of sequenced microbial genes. This community page discusses the progress made by the COMBREX project to address this important issue using both computational and experimental resources.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
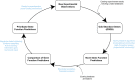
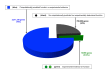
References
-
- Roberts RJ (2004) Identifying protein function—a call for community action. PLoS Biol 2: e42 doi:10.1371/journal.pbio.0020042 - DOI - PMC - PubMed
-
- Cohn D, Atlas L, Ladner R (1994) Improving generalization with active learning. Machine Learning 15: 201–221.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

