Mutations on the N-terminal edge of the DELSEED loop in either the α or β subunit of the mitochondrial F1-ATPase enhance ATP hydrolysis in the absence of the central γ rotor
- PMID: 24014764
- PMCID: PMC3837939
- DOI: 10.1128/EC.00177-13
Mutations on the N-terminal edge of the DELSEED loop in either the α or β subunit of the mitochondrial F1-ATPase enhance ATP hydrolysis in the absence of the central γ rotor
Abstract
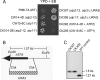
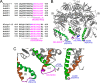
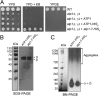
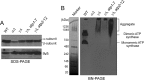
F(1)-ATPase is a rotary molecular machine with a subunit stoichiometry of α(3)β(3)γ(1)δ(1)ε(1). It has a robust ATP-hydrolyzing activity due to effective cooperativity between the three catalytic sites. It is believed that the central γ rotor dictates the sequential conformational changes to the catalytic sites in the α(3)β(3) core to achieve cooperativity. However, recent studies of the thermophilic Bacillus PS3 F(1)-ATPase have suggested that the α(3)β(3) core can intrinsically undergo unidirectional cooperative catalysis (T. Uchihashi et al., Science 333:755-758, 2011). The mechanism of this γ-independent ATP-hydrolyzing mode is unclear. Here, a unique genetic screen allowed us to identify specific mutations in the α and β subunits that stimulate ATP hydrolysis by the mitochondrial F(1)-ATPase in the absence of γ. We found that the F446I mutation in the α subunit and G419D mutation in the β subunit suppress cell death by the loss of mitochondrial DNA (ρ(o)) in a Kluyveromyces lactis mutant lacking γ. In organello ATPase assays showed that the mutant but not the wild-type γ-less F(1) complexes retained 21.7 to 44.6% of the native F(1)-ATPase activity. The γ-less F(1) subcomplex was assembled but was structurally and functionally labile in vitro. Phe446 in the α subunit and Gly419 in the β subunit are located on the N-terminal edge of the DELSEED loops in both subunits. Mutations in these two sites likely enhance the transmission of catalytically required conformational changes to an adjacent α or β subunit, thereby allowing robust ATP hydrolysis and cell survival under ρ(o) conditions. This work may help our understanding of the structural elements required for ATP hydrolysis by the α(3)β(3) subcomplex.
Figures



Similar articles
-
Inhibition of ATP hydrolysis by thermoalkaliphilic F1Fo-ATP synthase is controlled by the C terminus of the epsilon subunit.J Bacteriol. 2006 Jun;188(11):3796-804. doi: 10.1128/JB.00040-06. J Bacteriol. 2006. PMID: 16707672 Free PMC article.
-
Structures and interactions of proteins involved in the coupling function of the protonmotive F(o)F(1)-ATP synthase.Curr Protein Pept Sci. 2002 Aug;3(4):451-60. doi: 10.2174/1389203023380558. Curr Protein Pept Sci. 2002. PMID: 12370007 Review.
-
The F1-ATPase inhibitor Inh1 (IF1) affects suppression of mtDNA loss-lethality in Kluyveromyces lactis.FEMS Yeast Res. 2007 Aug;7(5):665-74. doi: 10.1111/j.1567-1364.2006.00201.x. Epub 2007 Feb 5. FEMS Yeast Res. 2007. PMID: 17286560
-
Absence of F1-ATPase activity in Kluyveromyces lactis lacking the epsilon subunit.Curr Genet. 2000 Jul;38(1):1-7. doi: 10.1007/s002940000127. Curr Genet. 2000. PMID: 10953875
-
Recent developments on structural and functional aspects of the F1 sector of H+-linked ATPases.Mol Cell Biochem. 1984;60(1):33-71. doi: 10.1007/BF00226299. Mol Cell Biochem. 1984. PMID: 6231469 Review.
Cited by
-
Interaction between γC87 and γR242 residues participates in energy coupling between catalysis and proton translocation in Escherichia coli ATP synthase.Biochim Biophys Acta Bioenerg. 2019 Aug 1;1860(8):679-687. doi: 10.1016/j.bbabio.2019.06.016. Epub 2019 Jun 25. Biochim Biophys Acta Bioenerg. 2019. PMID: 31251901 Free PMC article.
-
Computational Design of Inhibitors Targeting the Catalytic β Subunit of Escherichia coli FOF1-ATP Synthase.Antibiotics (Basel). 2022 Apr 22;11(5):557. doi: 10.3390/antibiotics11050557. Antibiotics (Basel). 2022. PMID: 35625201 Free PMC article.
-
Possible Involvement of F1F0-ATP synthase and Intracellular ATP in Keratinocyte Differentiation in normal skin and skin lesions.Sci Rep. 2017 Feb 17;7:42672. doi: 10.1038/srep42672. Sci Rep. 2017. PMID: 28209970 Free PMC article.
References
-
- Ephrussi B, Hottinguer H, Tavlitzki J. 1949. Action de l'acriflavine sur les levures. II. Etude génétique du mutant “petite colonie”. Ann. Inst. Pasteur (Paris) 76:419–442
-
- Bulder CJ. 1964. Lethality of the petite mutation in petite negative yeasts. Antonie Van Leeuwenhoek 30:442–454 - PubMed
-
- Bulder CJ. 1964. Induction of petite mutation and inhibition of synthesis of respiratory enzymes in various yeasts. Antonie Van Leeuwenhoek 30:1–9 - PubMed
-
- Faye G, Fukuhara H, Grandchamp C, Lazowska J, Michel F, Casey J, Getz GS, Locker J, Rabinowitz M, Bolotin-Fukuhara M, Coen D, Deutsch J, Dujon B, Netter P, Slonimski PP. 1973. Mitochondrial nucleic acids in the petite colonie mutants: deletions and repetition of genes. Biochimie 55:779–792 - PubMed
-
- Chen XJ, Clark-Walker GD. 2000. The petite mutation in yeasts: 50 years on. Int. Rev. Cytol. 194:197–238 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

