Phenol-soluble modulins and staphylococcal infection
- PMID: 24018382
- PMCID: PMC4780437
- DOI: 10.1038/nrmicro3110
Phenol-soluble modulins and staphylococcal infection
Erratum in
- Nat Rev Microbiol. 2013 Nov;11(11):814
Abstract
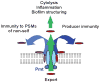
Staphylococcus aureus is an important human pathogen and a leading cause of death worldwide. Phenol-soluble modulins (PSMs) have recently emerged as a novel toxin family defining the virulence potential of highly aggressive S. aureus isolates. PSMs have multiple roles in staphylococcal pathogenesis, causing lysis of red and white blood cells, stimulating inflammatory responses and contributing to biofilm development and the dissemination of biofilm-associated infections. Moreover, the pronounced capacity of PSMs to kill human neutrophils after phagocytosis might explain failures in the development of anti-staphylococcal vaccines. Here, we discuss recent progress made in our understanding of the biochemical and genetic properties of PSMs and their role in S. aureus pathogenesis, and suggest potential avenues to target PSMs for the development of anti-staphylococcal drugs.
Figures



Similar articles
-
Phenol-soluble modulins--critical determinants of staphylococcal virulence.FEMS Microbiol Rev. 2014 Jul;38(4):698-719. doi: 10.1111/1574-6976.12057. Epub 2014 Jan 16. FEMS Microbiol Rev. 2014. PMID: 24372362 Free PMC article. Review.
-
Phenol-soluble modulins.Int J Med Microbiol. 2014 Mar;304(2):164-9. doi: 10.1016/j.ijmm.2013.11.019. Epub 2013 Dec 1. Int J Med Microbiol. 2014. PMID: 24447915 Free PMC article. Review.
-
Staphylococcus epidermidis strategies to avoid killing by human neutrophils.PLoS Pathog. 2010 Oct 7;6(10):e1001133. doi: 10.1371/journal.ppat.1001133. PLoS Pathog. 2010. PMID: 20949069 Free PMC article.
-
Role of Phenol-Soluble Modulins in Staphylococcus epidermidis Biofilm Formation and Infection of Indwelling Medical Devices.J Mol Biol. 2019 Jul 26;431(16):3015-3027. doi: 10.1016/j.jmb.2019.03.030. Epub 2019 Apr 5. J Mol Biol. 2019. PMID: 30954574 Free PMC article.
-
Phenol-soluble modulins: novel virulence-associated peptides of staphylococci.Future Microbiol. 2014;9(2):203-16. doi: 10.2217/fmb.13.153. Epub 2013 Dec 3. Future Microbiol. 2014. PMID: 24295365 Review.
Cited by
-
Comparative Analysis of the Microbiome across the Gut-Skin Axis in Atopic Dermatitis.Int J Mol Sci. 2021 Apr 19;22(8):4228. doi: 10.3390/ijms22084228. Int J Mol Sci. 2021. PMID: 33921772 Free PMC article. Review.
-
Molecular Characteristics and Pathogenicity of Staphylococcus aureus Exotoxins.Int J Mol Sci. 2023 Dec 28;25(1):395. doi: 10.3390/ijms25010395. Int J Mol Sci. 2023. PMID: 38203566 Free PMC article. Review.
-
Mucin-induced surface dispersal of Staphylococcus aureus and Staphylococcus epidermidis via quorum-sensing dependent and independent mechanisms.mBio. 2024 Aug 14;15(8):e0156224. doi: 10.1128/mbio.01562-24. Epub 2024 Jul 2. mBio. 2024. PMID: 38953351 Free PMC article.
-
Slightly acidic electrolyzed water inhibits inflammation induced by membrane vesicles of Staphylococcus aureus.Front Microbiol. 2024 Jan 12;14:1328055. doi: 10.3389/fmicb.2023.1328055. eCollection 2023. Front Microbiol. 2024. PMID: 38282743 Free PMC article.
-
Modulation of MRSA virulence gene expression by the wall teichoic acid enzyme TarO.Nat Commun. 2023 Mar 22;14(1):1594. doi: 10.1038/s41467-023-37310-5. Nat Commun. 2023. PMID: 36949052 Free PMC article.
References
-
- Wertheim HF, et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect Dis. 2005;5:751–62. - PubMed
-
- Lowy FD. Staphylococcus aureus infections. N Engl J Med. 1998;339:520–32. - PubMed
-
- Klevens RM, et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. Jama. 2007;298:1763–71. - PubMed
-
- Costerton JW, Stewart PS, Greenberg EP. Bacterial biofilms: a common cause of persistent infections. Science. 1999;284:1318–22. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

