The FBXO4 tumor suppressor functions as a barrier to BRAFV600E-dependent metastatic melanoma
- PMID: 24019069
- PMCID: PMC3838190
- DOI: 10.1128/MCB.00706-13
The FBXO4 tumor suppressor functions as a barrier to BRAFV600E-dependent metastatic melanoma
Abstract
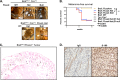
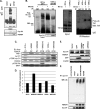
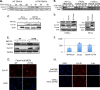
Cyclin D1-cyclin-dependent kinase 4/6 (CDK4/6) dysregulation is a major contributor to melanomagenesis. Clinical evidence has revealed that p16(INK4A), an allosteric inhibitor of CDK4/6, is inactivated in over half of human melanomas, and numerous animal models have demonstrated that p16(INK4A) deletion promotes melanoma. FBXO4, a specificity factor for the E3 ligase that directs timely cyclin D1 proteolysis, has not been studied in melanoma. We demonstrate that Fbxo4 deficiency induces Braf-driven melanoma and that this phenotype depends on cyclin D1 accumulation in mice, underscoring the importance of this ubiquitin ligase in tumor suppression. Furthermore, we have identified a substrate-binding mutation, FBXO4 I377M, that selectively disrupts cyclin D1 degradation while preserving proteolysis of the other known FBXO4 substrate, TRF1. The I377M mutation and Fbxo4 deficiency result in nuclear accumulation of cyclin D1, a key transforming neoplastic event. Collectively, these data provide evidence that FBXO4 dysfunction, as a mechanism for cyclin D1 overexpression, is a contributor to human malignancy.
Figures



Similar articles
-
FBXO4 loss facilitates carcinogen induced papilloma development in mice.Cancer Biol Ther. 2015;16(5):750-5. doi: 10.1080/15384047.2015.1026512. Cancer Biol Ther. 2015. PMID: 25801820 Free PMC article.
-
Increased cyclin D1 expression can mediate BRAF inhibitor resistance in BRAF V600E-mutated melanomas.Mol Cancer Ther. 2008 Sep;7(9):2876-83. doi: 10.1158/1535-7163.MCT-08-0431. Mol Cancer Ther. 2008. PMID: 18790768 Free PMC article.
-
Genetic reevaluation of the role of F-box proteins in cyclin D1 degradation.Mol Cell Biol. 2012 Feb;32(3):590-605. doi: 10.1128/MCB.06570-11. Epub 2011 Nov 28. Mol Cell Biol. 2012. PMID: 22124152 Free PMC article.
-
The E3 Ubiquitin Ligase Fbxo4 Functions as a Tumor Suppressor: Its Biological Importance and Therapeutic Perspectives.Cancers (Basel). 2022 Apr 25;14(9):2133. doi: 10.3390/cancers14092133. Cancers (Basel). 2022. PMID: 35565262 Free PMC article. Review.
-
SCF(Fbx4/alphaB-crystallin) E3 ligase: when one is not enough.Cell Cycle. 2008 Oct;7(19):2983-6. doi: 10.4161/cc.7.19.6775. Epub 2008 Oct 12. Cell Cycle. 2008. PMID: 18818515 Review.
Cited by
-
Prognostic values of F-box members in breast cancer: an online database analysis and literature review.Biosci Rep. 2019 Jan 3;39(1):BSR20180949. doi: 10.1042/BSR20180949. Print 2019 Jan 31. Biosci Rep. 2019. PMID: 30341246 Free PMC article. Review.
-
Cyclin D1, cancer progression, and opportunities in cancer treatment.J Mol Med (Berl). 2016 Dec;94(12):1313-1326. doi: 10.1007/s00109-016-1475-3. Epub 2016 Oct 2. J Mol Med (Berl). 2016. PMID: 27695879 Free PMC article. Review.
-
Ubiquitination in melanoma pathogenesis and treatment.Cancer Med. 2017 Jun;6(6):1362-1377. doi: 10.1002/cam4.1069. Epub 2017 May 23. Cancer Med. 2017. PMID: 28544818 Free PMC article. Review.
-
Tumor suppressor mediated ubiquitylation of hnRNPK is a barrier to oncogenic translation.Nat Commun. 2022 Nov 3;13(1):6614. doi: 10.1038/s41467-022-34402-6. Nat Commun. 2022. PMID: 36329064 Free PMC article.
-
Recent advances in SCF ubiquitin ligase complex: Clinical implications.Biochim Biophys Acta. 2016 Aug;1866(1):12-22. doi: 10.1016/j.bbcan.2016.05.001. Epub 2016 May 5. Biochim Biophys Acta. 2016. PMID: 27156687 Free PMC article. Review.
References
-
- Chin L. 2003. The genetics of malignant melanoma: lessons from mouse and man. Nat. Rev. Cancer 3:559–570 - PubMed
-
- Chin L, Merlino G, DePinho RA. 1998. Malignant melanoma: modern black plague and genetic black box. Genes Dev. 12:3467–3481 - PubMed
-
- Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA. 2002. Mutations of the BRAF gene in human cancer. Nature 417:949–954 - PubMed
-
- Pollock PM, Harper UL, Hansen KS, Yudt LM, Stark M, Robbins CM, Moses TY, Hostetter G, Wagner U, Kakareka J, Salem G, Pohida T, Heenan P, Duray P, Kallioniemi O, Hayward NK, Trent JM, Meltzer PS. 2003. High frequency of BRAF mutations in nevi. Nat. Genet. 33:19–20 - PubMed
-
- Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, Jouary T, Schadendorf D, Ribas A, O'Day SJ, Sosman JA, Kirkwood JM, Eggermont AM, Dreno B, Nolop K, Li J, Nelson B, Hou J, Lee RJ, Flaherty KT, McArthur GA. 2011. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N. Engl. J. Med. 364:2507–2516 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
