Histology-derived volumetric annotation of the human hippocampal subfields in postmortem MRI
- PMID: 24036353
- PMCID: PMC3864597
- DOI: 10.1016/j.neuroimage.2013.08.067
Histology-derived volumetric annotation of the human hippocampal subfields in postmortem MRI
Abstract
Recently, there has been a growing effort to analyze the morphometry of hippocampal subfields using both in vivo and postmortem magnetic resonance imaging (MRI). However, given that boundaries between subregions of the hippocampal formation (HF) are conventionally defined on the basis of microscopic features that often lack discernible signature in MRI, subfield delineation in MRI literature has largely relied on heuristic geometric rules, the validity of which with respect to the underlying anatomy is largely unknown. The development and evaluation of such rules are challenged by the limited availability of data linking MRI appearance to microscopic hippocampal anatomy, particularly in three dimensions (3D). The present paper, for the first time, demonstrates the feasibility of labeling hippocampal subfields in a high resolution volumetric MRI dataset based directly on microscopic features extracted from histology. It uses a combination of computational techniques and manual post-processing to map subfield boundaries from a stack of histology images (obtained with 200μm spacing and 5μm slice thickness; stained using the Kluver-Barrera method) onto a postmortem 9.4Tesla MRI scan of the intact, whole hippocampal formation acquired with 160μm isotropic resolution. The histology reconstruction procedure consists of sequential application of a graph-theoretic slice stacking algorithm that mitigates the effects of distorted slices, followed by iterative affine and diffeomorphic co-registration to postmortem MRI scans of approximately 1cm-thick tissue sub-blocks acquired with 200μm isotropic resolution. These 1cm blocks are subsequently co-registered to the MRI of the whole HF. Reconstruction accuracy is evaluated as the average displacement error between boundaries manually delineated in both the histology and MRI following the sequential stages of reconstruction. The methods presented and evaluated in this single-subject study can potentially be applied to multiple hippocampal tissue samples in order to construct a histologically informed MRI atlas of the hippocampal formation.
Keywords: Hippocampus; Histology; MRI; Postmortem; Reconstruction; Subfields.
© 2013 Elsevier Inc. All rights reserved.
Figures






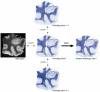








References
-
- Amaral D, Lavenex P. The Hippocampus Book. Oxford University Press; Cambridge: 2007.
-
- Amunts K, Kedo O, Kindler M, Pieperhoff P, Mohlberg H, Shah N, Habel U, Schneider F, Zilles K. Cytoarchitectonic mapping of the human amygdala, hippocampal region and entorhinal cortex: intersubject variability and probability maps. Anat Embryol. 2005;210(5-6):343–352. - PubMed
-
- Amunts K, Lepage C, Borgeat L, Mohlberg H, Dickscheid T, Rousseau M-, Bludau S, Bazin P-L, Lewis LB, Oros-Peusquens A-M. BigBrain: an ultrahigh-resolution 3D human brain model. Science. 2013;340(6139):14721475. - PubMed
-
- Apostolova LG, Dinov ID, Dutton RA, Hayashi KM, Toga AW, Cummings JL, Thompson PM. 3D comparison of hippocampal atrophy in amnestic mild cognitive impairment and Alzheimer’s disease. Brain. 2006;129(11):2867–2873. - PubMed
-
- Arganda-Carreras I, Sorzano COS, Thévenaz P, Muñoz Barrutia A, Kybic J, Marabini R, Carazo JM, Ortiz-de Solorzano C. Non-rigid consistent registration of 2D image sequences. Phys Med Biol. 2010;55(20):6215–6242. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

