Transcriptome profiling of Gossypium barbadense inoculated with Verticillium dahliae provides a resource for cotton improvement
- PMID: 24053558
- PMCID: PMC3849602
- DOI: 10.1186/1471-2164-14-637
Transcriptome profiling of Gossypium barbadense inoculated with Verticillium dahliae provides a resource for cotton improvement
Abstract
Background: Verticillium wilt, caused by the fungal pathogen Verticillium dahliae, is the most severe disease in cotton (Gossypium spp.), causing great lint losses worldwide. Disease management could be achieved in the field if genetically improved, resistant plants were used. However, the interaction between V. dahliae and cotton is a complicated process, and its molecular mechanism remains obscure. To understand better the defense response to this pathogen as a means for obtaining more tolerant cultivars, we monitored the transcriptome profiles of roots from resistant plants of G. barbadense cv. Pima90-53 that were challenged with V. dahliae.
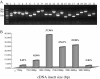
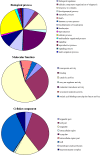
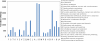
Results: In all, 46,192 high-quality expressed sequence tags (ESTs) were generated from a full-length cDNA library of G. barbadense. They were clustered and assembled into 23126 unigenes that comprised 2661 contigs and 20465 singletons. Those unigenes were assigned Gene Ontology terms and mapped to 289 KEGG pathways. A total of 3027 unigenes were found to be homologous to known defense-related genes in other plants. They were assigned to the functional classification of plant-pathogen interactions, including disease defenses and signal transduction. The branch of "SA→NPR1→TGA→PR-1→Disease resistance" was first discovered in the interaction of cotton-V. dahliae, indicating that this wilt process includes both biotrophic and necrotrophic stages. In all, 4936 genes coding for putative transcription factors (TF) were identified in our library. The most abundant TF family was the NAC group (527), followed by G2-like (440), MYB (372), BHLH (331), bZIP (271) ERF, C3H, and WRKY. We also analyzed the expression of genes involved in pathogen-associated molecular pattern (PAMP) recognition, the activation of effector-triggered immunity, TFs, and hormone biosynthesis, as well as genes that are pathogenesis-related, or have roles in signaling/regulatory functions and cell wall modification. Their differential expression patterns were compared among mock-/inoculated- and resistant/susceptible cotton. Our results suggest that the cotton defense response has significant transcriptional complexity and that large accumulations of defense-related transcripts may contribute to V. dahliae resistance in cotton. Therefore, these data provide a resource for cotton improvement through molecular breeding approaches.
Conclusions: This study generated a substantial amount of cotton transcript sequences that are related to defense responses against V. dahliae. These genomics resources and knowledge of important related genes contribute to our understanding of host-pathogen interactions and the defense mechanisms utilized by G. barbadense, a non-model plant system. These tools can be applied in establishing a modern breeding program that uses marker-assisted selections and oligonucleotide arrays to identify candidate genes that can be linked to valuable agronomic traits in cotton, including disease resistance.
Figures

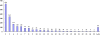

Similar articles
-
Transcriptome Analysis of a Cotton Cultivar Provides Insights into the Differentially Expressed Genes Underlying Heightened Resistance to the Devastating Verticillium Wilt.Cells. 2021 Oct 30;10(11):2961. doi: 10.3390/cells10112961. Cells. 2021. PMID: 34831184 Free PMC article.
-
Dynamic characteristics and functional analysis provide new insights into long non-coding RNA responsive to Verticillium dahliae infection in Gossypium hirsutum.BMC Plant Biol. 2021 Feb 1;21(1):68. doi: 10.1186/s12870-021-02835-8. BMC Plant Biol. 2021. PMID: 33526028 Free PMC article.
-
Overexpression of 3-deoxy-7-phosphoheptulonate synthase gene from Gossypium hirsutum enhances Arabidopsis resistance to Verticillium wilt.Plant Cell Rep. 2015 Aug;34(8):1429-41. doi: 10.1007/s00299-015-1798-5. Epub 2015 May 1. Plant Cell Rep. 2015. PMID: 25929795
-
Insights to Gossypium defense response against Verticillium dahliae: the Cotton Cancer.Funct Integr Genomics. 2023 May 1;23(2):142. doi: 10.1007/s10142-023-01065-5. Funct Integr Genomics. 2023. PMID: 37121989 Review.
-
Physiological and molecular mechanism of defense in cotton against Verticillium dahliae.Plant Physiol Biochem. 2018 Apr;125:193-204. doi: 10.1016/j.plaphy.2018.02.011. Epub 2018 Feb 13. Plant Physiol Biochem. 2018. PMID: 29462745 Review.
Cited by
-
Identification of Multiple Stress Responsive Genes by Sequencing a Normalized cDNA Library from Sea-Land Cotton (Gossypium barbadense L.).PLoS One. 2016 Mar 31;11(3):e0152927. doi: 10.1371/journal.pone.0152927. eCollection 2016. PLoS One. 2016. PMID: 27031331 Free PMC article.
-
Transcriptome analysis reveals differential transcription in tomato (Solanum lycopersicum) following inoculation with Ralstonia solanacearum.Sci Rep. 2022 Dec 22;12(1):22137. doi: 10.1038/s41598-022-26693-y. Sci Rep. 2022. PMID: 36550145 Free PMC article.
-
GhWRKY70D13 Regulates Resistance to Verticillium dahliae in Cotton Through the Ethylene and Jasmonic Acid Signaling Pathways.Front Plant Sci. 2020 Feb 25;11:69. doi: 10.3389/fpls.2020.00069. eCollection 2020. Front Plant Sci. 2020. PMID: 32158454 Free PMC article.
-
Genome-wide analysis of PRR gene family uncovers their roles in circadian rhythmic changes and response to drought stress in Gossypium hirsutum L.PeerJ. 2020 Sep 25;8:e9936. doi: 10.7717/peerj.9936. eCollection 2020. PeerJ. 2020. PMID: 33033660 Free PMC article.
-
Elucidating the molecular responses of apple rootstock resistant to ARD pathogens: challenges and opportunities for development of genomics-assisted breeding tools.Hortic Res. 2014 Aug 20;1:14043. doi: 10.1038/hortres.2014.43. eCollection 2014. Hortic Res. 2014. PMID: 26504547 Free PMC article. Review.
References
-
- Sink K, Grey WE. A root-injection method to assess Verticillium wilt resistance of peppermint (Mentha × piperita L) and its use in identifying resistant somaclones of cv Black Mitcham. Euphytica. 1999;106:223–230. doi: 10.1023/A:1003591908308. - DOI
-
- Kamal ME. Integrated control of Verticillium wilt of cotton. Plant Dis. 1985;69:1025–1032. doi: 10.1094/PD-69-1025. - DOI
-
- Wang XF, Wang YX, Zhang GY, Ma ZY. An integrated breeding technology for accelerating generation advancement and trait introgression in cotton. Plant Breed. 2011;130:569–573. doi: 10.1111/j.1439-0523.2011.01868.x. - DOI
-
- Cai YF, He XH, Mo JC, Sun Q, Yang JP, Liu JG. Molecular research and genetic engineering of resistance to Verticillium wilt in cotton. Afr J Biotechnol. 2009;8:7363–7372.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

