Antigen availability determines CD8⁺ T cell-dendritic cell interaction kinetics and memory fate decisions
- PMID: 24054328
- PMCID: PMC3914670
- DOI: 10.1016/j.immuni.2013.08.034
Antigen availability determines CD8⁺ T cell-dendritic cell interaction kinetics and memory fate decisions
Abstract
T cells are activated by antigen (Ag)-bearing dendritic cells (DCs) in lymph nodes in three phases. The duration of the initial phase of transient, serial DC-T cell interactions is inversely correlated with Ag dose. The second phase, characterized by stable DC-T cell contacts, is believed to be necessary for full-fledged T cell activation. Here we have shown that this is not the case. CD8⁺ T cells interacting with DCs presenting low-dose, short-lived Ag did not transition to phase 2, whereas higher Ag dose yielded phase 2 transition. Both antigenic constellations promoted T cell proliferation and effector differentiation but yielded different transcriptome signatures at 12 hr and 24 hr. T cells that experienced phase 2 developed long-lived memory, whereas conditions without stable contacts yielded immunological amnesia. Thus, T cells make fate decisions within hours after Ag exposure, resulting in long-term memory or abortive effector responses, correlating with T cell-DCs interaction kinetics.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures
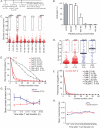

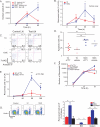
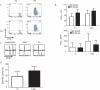


References
-
- Achour A, Michaelsson J, Harris RA, Odeberg J, Grufman P, Sandberg JK, Levitsky V, Karre K, Sandalova T, Schneider G. A structural basis for LCMV immune evasion: subversion of H-2D(b) and H-2K(b) presentation of gp33 revealed by comparative crystal structure analyses. Immunity. 2002;17:757–768. - PubMed
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

