Transcriptional mechanisms underlying sensitization of peripheral sensory neurons by granulocyte-/granulocyte-macrophage colony stimulating factors
- PMID: 24067145
- PMCID: PMC3852053
- DOI: 10.1186/1744-8069-9-48
Transcriptional mechanisms underlying sensitization of peripheral sensory neurons by granulocyte-/granulocyte-macrophage colony stimulating factors
Abstract
Background: Cancer-associated pain is a major cause of poor quality of life in cancer patients and is frequently resistant to conventional therapy. Recent studies indicate that some hematopoietic growth factors, namely granulocyte macrophage colony stimulating factor (GMCSF) and granulocyte colony stimulating factor (GCSF), are abundantly released in the tumor microenvironment and play a key role in regulating tumor-nerve interactions and tumor-associated pain by activating receptors on dorsal root ganglion (DRG) neurons. Moreover, these hematopoietic factors have been highly implicated in postsurgical pain, inflammatory pain and osteoarthritic pain. However, the molecular mechanisms via which G-/GMCSF bring about nociceptive sensitization and elicit pain are not known.
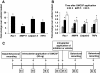
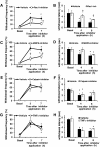
Results: In order to elucidate G-/GMCSF mediated transcriptional changes in the sensory neurons, we performed a comprehensive, genome-wide analysis of changes in the transcriptome of DRG neurons brought about by exposure to GMCSF or GCSF. We present complete information on regulated genes and validated profiling analyses and report novel regulatory networks and interaction maps revealed by detailed bioinformatics analyses. Amongst these, we validate calpain 2, matrix metalloproteinase 9 (MMP9) and a RhoGTPase Rac1 as well as Tumor necrosis factor alpha (TNFα) as transcriptional targets of G-/GMCSF and demonstrate the importance of MMP9 and Rac1 in GMCSF-induced nociceptor sensitization.
Conclusion: With integrative approach of bioinformatics, in vivo pharmacology and behavioral analyses, our results not only indicate that transcriptional control by G-/GMCSF signaling regulates a variety of established pain modulators, but also uncover a large number of novel targets, paving the way for translational analyses in the context of pain disorders.
Figures



References
-
- Stosser S, Agarwal N, Tappe-Theodor A, Yanagisawa M, Kuner R. Dissecting the functional significance of endothelin A receptors in peripheral nociceptors in vivo via conditional gene deletion. Pain. 2009;148:206–214. - PubMed
-
- Mantyh PW. Cancer pain and its impact on diagnosis, survival and quality of life. Nat Rev Neurosci. 2006;7:797–809. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

