Phenotypic and genetic consequences of protein damage
- PMID: 24068972
- PMCID: PMC3778015
- DOI: 10.1371/journal.pgen.1003810
Phenotypic and genetic consequences of protein damage
Abstract
Although the genome contains all the information necessary for maintenance and perpetuation of life, it is the proteome that repairs, duplicates and expresses the genome and actually performs most cellular functions. Here we reveal strong phenotypes of physiological oxidative proteome damage at the functional and genomic levels. Genome-wide mutations rates and biosynthetic capacity were monitored in real time, in single Escherichia coli cells with identical levels of reactive oxygen species and oxidative DNA damage, but with different levels of irreversible oxidative proteome damage (carbonylation). Increased protein carbonylation correlates with a mutator phenotype, whereas reducing it below wild type level produces an anti-mutator phenotype identifying proteome damage as the leading cause of spontaneous mutations. Proteome oxidation elevates also UV-light induced mutagenesis and impairs cellular biosynthesis. In conclusion, protein damage reduces the efficacy and precision of vital cellular processes resulting in high mutation rates and functional degeneracy akin to cellular aging.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
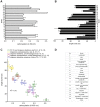
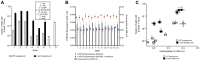



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

